HDAC2 (UniProt/Swiss-Prot entry Q92769) belongs to the family of histone deacetylases which catalyse the deacetylation of lysine residues on the N-terminal part of the core histones (H2A, H2B, H3 and H4). Acetylation and deacetylation of these highly conserved lysine residues is important for the control of gene expression and HDAC2 activity is associated with gene repression. Histone deacetylation is established by the formation of large multiprotein complexes. Thereby, HDAC2 associates with many different proteins, including the mammalian zinc-finger transcription factor YY1. HDAC2 thus plays an important role in transcriptional regulation, cell cycle progression and developmental events.
HDAC2 monoclonal antibody
| Lot | 001 |
|---|---|
| Concentration | 2.0 µg/µl |
| Species reactivity | Human |
| Type | Monoclonal |
| Purity | Protein A purified |
| Host | Mouse |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Applications | Suggested dilution | References |
|---|---|---|
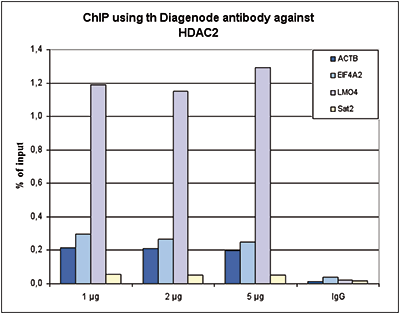
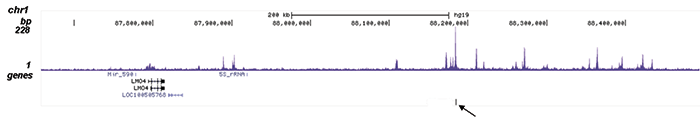
| ChIP/ChIP-seq * | 1 μg/IP | Fig 1, 2 |
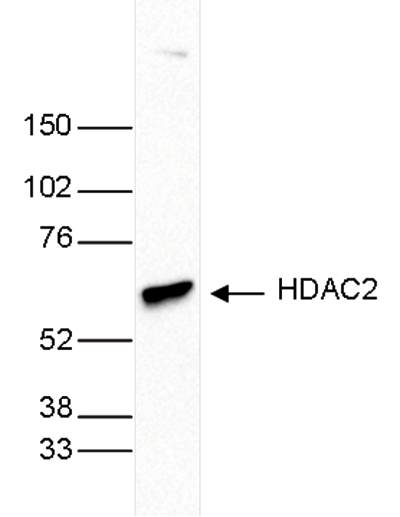
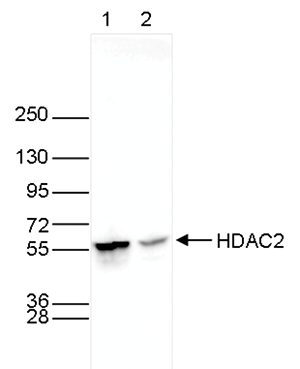
| Western Blotting | 1:1,000 | Fig 3, 4 |
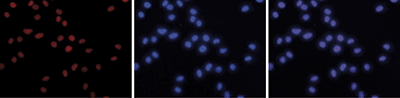
| Immunofluorescence | 1:500 | Fig 5 |
* Please note that the optimal antibody amount per IP should be determined by the end-user. We recommend testing 1-5 μg per IP.