We were very happy with the method. It gave good results in the end, and required much smaller samples than we need to reliably perform conventional ChIP-seq.
In our view, the main advantages of the ChIPmentation kit compared to our conventional ChIP-seq protocol are (most important first):
- smaller sample requirement,
- simpler workflow with less that can go wrong,
- slightly higher resolution and signal: noise ratio.
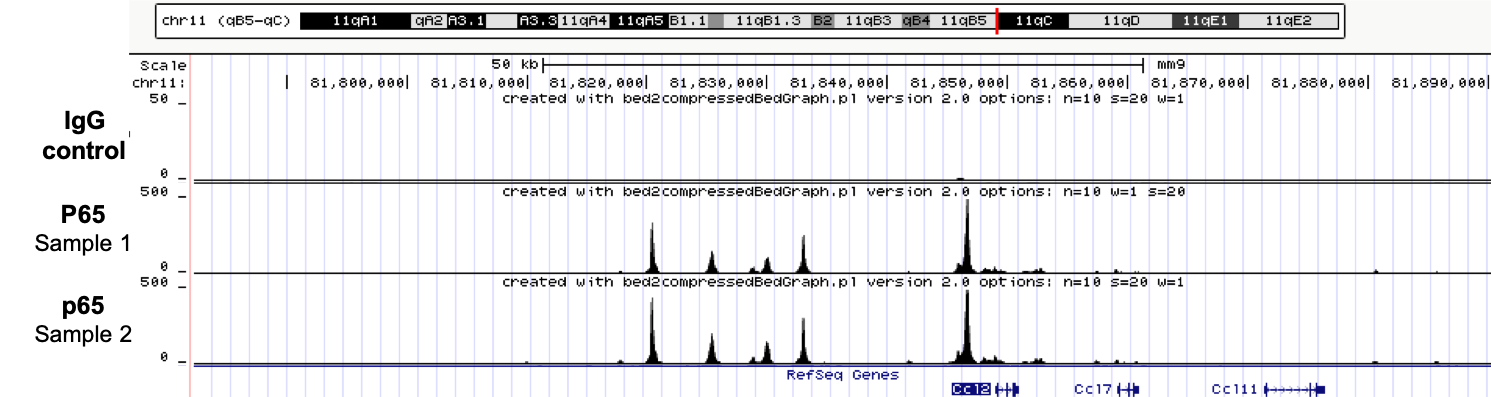
ChIPmentation sequencing profiles for p65. Chromatin preparation and immunoprecipitation have been performed on stimulated NIH3T3 cells using the iDeal ChIP-seq kit for TFs (Cat. No. C01010055). Chromatin from 4,000,000 cells was used for the immunoprecipitation in combination with either anti-p65 antibody or IgG. The library preparation was performed with the TAG Kit for ChIPmentation (Cat. No. C01011030) and 24 SI for ChIPmentation (Cat. No. C01011031).
TAG Kit for ChIPmentation
Tagmentationは、酵素(トランスポザーゼ)がDNAを切断し、DNAフラグメントの末端にシーケンスアダプターを1ステップで組み込む反応を言います。 DiagenodeのChIPmentationテクノロジーでは、クロマチン免疫沈降とタグ付けを1つの合理化されたワークフローで組み合わせ、Tagmentationがクロマチンで直接行われます。 TAG Kit for ChIPmentationは、独自の検証済みChIPプロトコールを使用してChIPmentation専用として開発されました。 TAG Kit for ChIPmentationには、Tagmentationベースのライブラリー調製用の試薬が含まれており、磁気ビーズに基づき、どんなChIPプロトコールとも互換性があります。 多重化のインデックスは個別に購入頂く必要があり、参照としてご利用頂けます。:24 SI for ChIPmentation、Cat。 番号C01011031
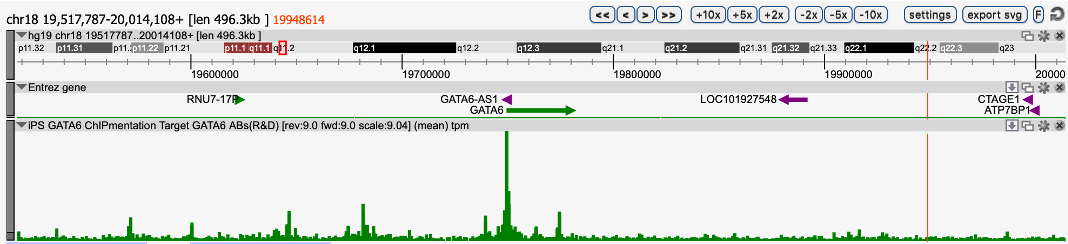
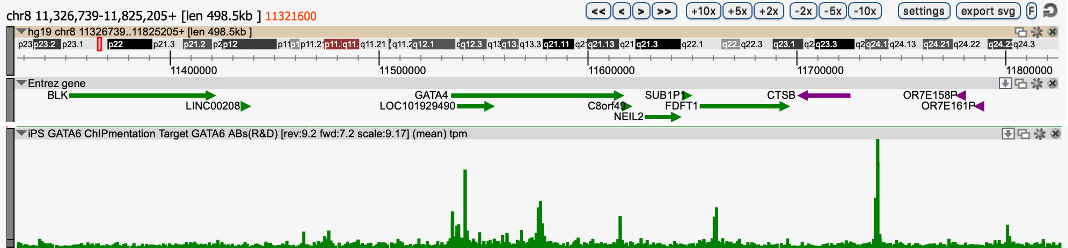
TESTIMONIALOne of our issues was that we could obtain only a limited number of cells, which is not enough for canonical ChIP-seq protocols. To solve this issue, we used the Diagenode ChIPmentation solution composed of iDeal ChIP-seq Kit for Transcription Factor, TAG Kit for ChIPmentation, and 24 SI for ChIPmentation. We performed ChIPmentation with IP-Star automated system for GATA6 in 2 million GATA6-overxpressing iPS cells. The result showed clear signal/noise ratio and was highly reproducible. This solution also worked in vitro differentiated definitive endoderm cells (data not shown).
Region 1
Region 2
Figure 1. ChIPmentation sequencing profiles for Gata6
Chromatin preparation and immunoprecipitation have been performed on hiPSCs (human induced Pluripotent Stem Cells) overexpressing Gata6 using the iDeal ChIP-seq kit for TFs (Cat. No. C01010055). Chromatin from 2,000,000 cells was used for the immunoprecipitation in combination with either anti-GATA6 antibody. The library preparation was performed with the TAG Kit for ChIPmentation (Cat. No. C01011030) and 24 SI for ChIPmentation (Cat. No. C01011031).