Notice (8): Undefined variable: solution_of_interest [APP/View/Products/view.ctp, line 755]Code Context<!-- BEGIN: REQUEST_FORM MODAL -->
<div id="request_formModal" class="reveal-modal medium" data-reveal aria-labelledby="modalTitle" aria-hidden="true" role="dialog">
<?= $this->element('Forms/simple_form', array('solution_of_interest' => $solution_of_interest, 'header' => $header, 'message' => $message, 'campaign_id' => $campaign_id)) ?>
$viewFile = '/home/website-server/www/app/View/Products/view.ctp'
$dataForView = array(
'language' => 'en',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'product' => array(
'Product' => array(
'id' => '3027',
'antibody_id' => null,
'name' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'description' => '<p>Diagenode’s <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns">MicroPlex Library Preparation Kits v3</a> have been extensively validated for ChIP-seq library prep from ChIP-derived DNA. The kit MicroPlex v3 has to be purchased with the compatible set of dual indexes. This set of dual indexes allows for multiplexing up to 96 samples.</p>
<p>Check all available sets of dual indexes:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong></li>
<li><strong>Validated with the </strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">IP-Star Compact Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
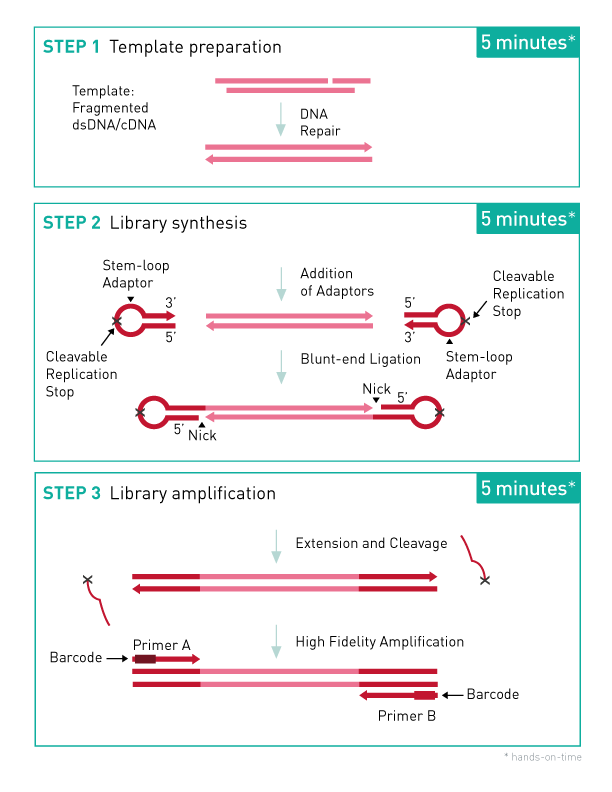
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>',
'label2' => '',
'info2' => '',
'label3' => '',
'info3' => '',
'format' => '96 rxns',
'catalog_number' => 'C05010005',
'old_catalog_number' => '',
'sf_code' => 'C05010005-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1355',
'price_USD' => '1300',
'price_GBP' => '1235',
'price_JPY' => '222015',
'price_CNY' => '',
'price_AUD' => '3250',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => '96-dual-indexes-for-microplex-kit-v3-set-2',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'modified' => '2020-10-23 17:11:41',
'created' => '2019-07-16 15:48:28',
'locale' => 'eng'
),
'Antibody' => array(
'host' => '*****',
'id' => null,
'name' => null,
'description' => null,
'clonality' => null,
'isotype' => null,
'lot' => null,
'concentration' => null,
'reactivity' => null,
'type' => null,
'purity' => null,
'classification' => null,
'application_table' => null,
'storage_conditions' => null,
'storage_buffer' => null,
'precautions' => null,
'uniprot_acc' => null,
'slug' => null,
'meta_keywords' => null,
'meta_description' => null,
'modified' => null,
'created' => null,
'select_label' => null
),
'Slave' => array(),
'Group' => array(),
'Related' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Application' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Category' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Document' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Feature' => array(),
'Image' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Promotion' => array(),
'Protocol' => array(),
'Publication' => array(),
'Testimonial' => array(),
'Area' => array(),
'SafetySheet' => array()
)
)
$language = 'en'
$meta_keywords = ''
$meta_description = '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns'
$meta_title = '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns'
$product = array(
'Product' => array(
'id' => '3027',
'antibody_id' => null,
'name' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'description' => '<p>Diagenode’s <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns">MicroPlex Library Preparation Kits v3</a> have been extensively validated for ChIP-seq library prep from ChIP-derived DNA. The kit MicroPlex v3 has to be purchased with the compatible set of dual indexes. This set of dual indexes allows for multiplexing up to 96 samples.</p>
<p>Check all available sets of dual indexes:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong></li>
<li><strong>Validated with the </strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">IP-Star Compact Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>',
'label2' => '',
'info2' => '',
'label3' => '',
'info3' => '',
'format' => '96 rxns',
'catalog_number' => 'C05010005',
'old_catalog_number' => '',
'sf_code' => 'C05010005-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1355',
'price_USD' => '1300',
'price_GBP' => '1235',
'price_JPY' => '222015',
'price_CNY' => '',
'price_AUD' => '3250',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => '96-dual-indexes-for-microplex-kit-v3-set-2',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'modified' => '2020-10-23 17:11:41',
'created' => '2019-07-16 15:48:28',
'locale' => 'eng'
),
'Antibody' => array(
'host' => '*****',
'id' => null,
'name' => null,
'description' => null,
'clonality' => null,
'isotype' => null,
'lot' => null,
'concentration' => null,
'reactivity' => null,
'type' => null,
'purity' => null,
'classification' => null,
'application_table' => null,
'storage_conditions' => null,
'storage_buffer' => null,
'precautions' => null,
'uniprot_acc' => null,
'slug' => null,
'meta_keywords' => null,
'meta_description' => null,
'modified' => null,
'created' => null,
'select_label' => null
),
'Slave' => array(),
'Group' => array(),
'Related' => array(
(int) 0 => array(
'id' => '3032',
'antibody_id' => null,
'name' => 'MicroPlex Library Preparation Kit v3 /48 rxns',
'description' => '<p><a href="https://www.diagenode.com/files/products/kits/Microplex-library-prep-v3.pdf"><img src="https://www.diagenode.com/img/buttons/bt-manual.png" /></a></p>
<p>Diagenode’s <strong>MicroPlex Library Preparation Kits v3</strong> have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg. The kit MicroPlex v3 includes all buffers and enzymes necessary for the library preparation. For flexibility of the choice different formats of compatible primer indexes are available separately:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2">C05010005 - 96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p style="padding-left: 30px;">NEW! Unique dual indexes :</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1">C05010008 - 24 UDI for MicroPlex Kit v3 - Set I</a></li>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2">C05010009 - 24 UDI for MicroPlex Kit v3 - Set II</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong> with 24 dual indexes (8 nt)</li>
<li><strong>Validated with the IP-<a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">Star<sup>®</sup></a></strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit"> Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img alt="MicroPlex Library Preparation Kit v3 /48 rxns" src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label2' => '',
'info2' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label3' => '',
'info3' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'format' => '48 rxns',
'catalog_number' => 'C05010001',
'old_catalog_number' => '',
'sf_code' => 'C05010001-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1750',
'price_USD' => '1555',
'price_GBP' => '1555',
'price_JPY' => '286740',
'price_CNY' => '',
'price_AUD' => '3888',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => 'microplex-lib-prep-kit-v3-48-rxns',
'meta_title' => 'MicroPlex Library Preparation Kit v3 /48 rxns | Diagenode',
'meta_keywords' => 'MicroPlex Library Preparation Kit',
'meta_description' => 'MicroPlex Library Preparation Kits v3 have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg.',
'modified' => '2023-04-06 12:08:35',
'created' => '2019-07-16 16:03:58',
'ProductsRelated' => array(
[maximum depth reached]
),
'Image' => array(
[maximum depth reached]
)
)
),
'Application' => array(
(int) 0 => array(
'id' => '14',
'position' => '10',
'parent_id' => '3',
'name' => 'DNA/RNA library preparation',
'description' => '<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p><span style="font-weight: 400;">Most of the major next-generation sequencing platforms require ligation of specific adaptor oligos to </span><a href="../applications/dna-rna-shearing"><span style="font-weight: 400;">fragmented DNA or RNA</span></a><span style="font-weight: 400;"> prior to sequencing</span></p>
<p><span style="font-weight: 400;">After input DNA has been fragmented, it is end-repaired and blunt-ended</span><span style="font-weight: 400;">. The next step is a A-tailing in which dAMP is added to the 3´ end of the blunt phosphorylated DNA fragments to prevent concatemerization and to allow the ligation of adaptors with complementary dT overhangs. In addition, barcoded adapters can be incorporated to facilitate multiplexing prior to or during amplification.</span></p>
<center><img src="https://www.diagenode.com/img/categories/library-prep/flux.png" /></center>
<p><span style="font-weight: 400;">Diagenode offers a comprehensive product portfolio for library preparation:<br /></span></p>
<strong><a href="https://www.diagenode.com/en/categories/Library-preparation-for-RNA-seq">D-Plex RNA-seq Library Preparation Kits</a></strong><br />
<p><span style="font-weight: 400;">Diagenode’s new RNA-sequencing solutions utilize the innovative c</span><span style="font-weight: 400;">apture and a</span><span style="font-weight: 400;">mplification by t</span><span style="font-weight: 400;">ailing and s</span><span style="font-weight: 400;">witching”</span><span style="font-weight: 400;">, a ligation-free method to produce DNA libraries for next generation sequencing from low input amounts of RNA. </span><span style="font-weight: 400;"></span><a href="../categories/Library-preparation-for-RNA-seq">Learn more</a></p>
<strong><a href="../categories/library-preparation-for-ChIP-seq">ChIP-seq and DNA sequencing library preparation solutions</a></strong><br />
<p><span style="font-weight: 400;">Our kits have been optimized for DNA library preparation used for next generation sequencing for a wide range of inputs. Using a simple three-step protocols, our</span><a href="http://www.diagenode.com/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns"><span style="font-weight: 400;"> </span></a><span style="font-weight: 400;">kits are an optimal choice for library preparation from DNA inputs down to 50 pg. </span><a href="../categories/library-preparation-for-ChIP-seq">Learn more</a></p>
<a href="../p/bioruptor-pico-sonication-device"><span style="font-weight: 400;"></span><strong>Bioruptor Pico - short fragments</strong></a><a href="../categories/library-preparation-for-ChIP-seq-and-DNA-sequencing"><span style="font-weight: 400;"></span></a><br />
<p><span style="font-weight: 400;"></span><span style="font-weight: 400;">Our well-cited Bioruptor Pico is the shearing device of choice for chromatin and DNA fragmentation. Obtain uniform and tight fragment distributions between 150bp -2kb. </span><a href="../p/bioruptor-pico-sonication-device">Learn more</a></p>
<strong><a href="../p/megaruptor2-1-unit"><span href="../p/bioruptor-pico-sonication-device">Megaruptor</span>® - long fragments</a></strong><a href="../p/bioruptor-pico-sonication-device"><span style="font-weight: 400;"></span></a><a href="../categories/library-preparation-for-ChIP-seq-and-DNA-sequencing"><span style="font-weight: 400;"></span></a><br />
<p><span style="font-weight: 400;"></span><span style="font-weight: 400;">The Megaruptor is designed to shear DNA from 3kb-75kb for long-read sequencing. <a href="../p/megaruptor2-1-unit">Learn more</a></span></p>
<span href="../p/bioruptor-pico-sonication-device"></span><span style="font-weight: 400;"></span></div>
</div>',
'in_footer' => false,
'in_menu' => true,
'online' => true,
'tabular' => true,
'slug' => 'library-preparation',
'meta_keywords' => 'Library preparation,Next Generation Sequencing,DNA fragments,MicroPlex Library,iDeal Library',
'meta_description' => 'Diagenode offers a comprehensive product portfolio for library preparation such as CATS RNA-seq Library Preparation Kits,ChIP-seq and DNA sequencing library preparation solutions',
'meta_title' => 'Library preparation for Next Generation Sequencing (NGS) | Diagenode',
'modified' => '2021-03-10 03:27:16',
'created' => '2014-08-16 07:47:56',
'ProductsApplication' => array(
[maximum depth reached]
)
)
),
'Category' => array(
(int) 0 => array(
'id' => '124',
'position' => '1',
'parent_id' => '15',
'name' => 'Library preparation for ChIP-seq',
'description' => '<div class="row">
<div class="large-12 columns text-justify">
<p>Library preparation following ChIP can be challenging due to the limited amount of DNA recovered after ChIP. Diagenode has developed the optimal solutions for ChIP-seq using two different approaches: the ligation-based library preparation on purified DNA or the tagmentation-based ChIPmentation.</p>
</div>
</div>
<div class="row extra-spaced">
<div class="large-12 columns"><center><a href="https://www.diagenode.com/en/pages/form-microplex-promo" target="_blank"></a></center></div>
</div>
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div id="portal" class="main-portal">
<div class="portal-inner"><nav class="portal-nav">
<ul data-tab="" class="tips-menu">
<li><a href="#panel1" class="tips portal button">Ligation-based library prep</a></li>
<li><a href="#panel2" class="tips portal button">ChIPmentation</a></li>
<li><a href="#panel3" class="tips portal button">Kit choice guide</a></li>
<li><a href="#panel4" class="tips portal button">Resources</a></li>
<li><a href="#panel5" class="tips portal button">FAQs</a></li>
</ul>
</nav></div>
</div>
<div class="tabs-content">
<div class="content active" id="panel1">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v5" style="color: #13b29c;"><i class="fa fa-caret-right"></i> Standard input library prep</a>
<div id="v5" class="content">
<div class="small-12 medium-12 large-12 columns">
<p>The <strong>iDeal Library Preparation Kit</strong> reliably converts DNA into indexed libraries for next-generation sequencing, with input amounts down to <strong>5 ng</strong>. Our kit offers a simple and fast workflow, high yields, and ready-to-sequence DNA on the Illumina platform.</p>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> <strong>Sample</strong>: Fragmented dsDNA</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Input</strong>: 5 ng – 1 µg</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Fast protocol</strong>: 3 hours</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Easy processing</strong>: 3 steps</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Indexing</strong>: single indexes for multiplexing up to 24 samples</li>
<li><i class="fa fa-arrow-circle-right"></i> Manual and automated protocols available</li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<ul class="square">
<li>MeDIP-seq library prep</li>
<li>Genomic DNA sequencing</li>
<li>High input ChIP-seq</li>
</ul>
</div>
<div class="extra-spaced">
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010020</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" style="color: #b21329;" target="_blank">iDeal Library Preparation Kit x24 (incl. Index Primer Set 1)</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010021</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" style="color: #b21329;" target="_blank">Index Primer Set 2 (iDeal Lib. Prep Kit x24)</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
</li>
</ul>
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v4" style="color: #13b29c;"><i class="fa fa-caret-right"></i> Low input library prep</a>
<div id="v4" class="content active"><center><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" target="_blank"><img src="https://www.diagenode.com/img/banners/banner-microplex-v3-580.jpg" class="extra-spaced" /></a></center>
<div align="center"><a href="https://www.diagenode.com/pages/form-microplex3" class="center alert radius button extra-spaced"><i class="fa fa-info"></i> Contact us</a></div>
<div class="extra-spaced">
<p>Diagenode’s <strong>MicroPlex Library Preparation kits</strong> have been extensively validated for ChIP-seq samples. Generated libraries are compatible with single-end or paired-end sequencing. MicroPlex chemistry (using stem-loop adapters ) is specifically developed and optimized to generate DNA libraries with high molecular complexity from the lowest input amounts. Only <strong>50 pg to 50 ng</strong> of fragmented double-stranded DNA is required for library preparation. The entire <strong>three-step workflow</strong> takes place in a <strong>single tube</strong> or well in about <strong>2 hours</strong>. No intermediate purification steps and no sample transfers are necessary to prevent handling errors and loss of valuable samples.</p>
</div>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> <strong>Sample</strong>: Fragmented dsDNA</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Low input</strong>: 50 pg – 50 ng</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Fast protocol</strong>: 2 hours</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Easy processing</strong>: 3 steps in 1 tube</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>No intermediate purification</strong></li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
<li><i class="fa fa-arrow-circle-right"></i> Manual and automated protocols available</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<ul class="square">
<li>ChIP-seq library prep from ChIP-derived DNA</li>
<li>Low input DNA sequencing</li>
</ul>
</div>
<h2>Two versions are available:</h2>
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v2" style="color: #13b29c;"><i class="fa fa-caret-right"></i> MicroPlex Library Preparation Kit v2 with single indexes</a>
<div id="v2" class="content">
<p>The MicroPlex Library Preparation Kit v2 contains all necessary reagents including single indexes for multiplexing up to 48 samples using single barcoding.</p>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010012</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v2 (12 indexes)</a></td>
<td class="format">12 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</li>
<li class="accordion-navigation"><a href="#v3" style="color: #13b29c;"><i class="fa fa-caret-right"></i> MicroPlex Library Preparation Kit v3 with dual indexes <strong><span class="diacol">NEW!</span></strong></a>
<div id="v3" class="content active">
<p>In this version the library preparation reagents and the dual indexes are available separately allowing for the flexibility choosing the number of indexes. MicroPlex v3 has multiplexing capacities up to 384 samples.</p>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010001</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v3 /48 rxns</a></td>
<td class="format">48 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010002</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v3 /96 rxns</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
<h4>DUAL INDEXES</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010003</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" style="color: #b21329;" target="_blank">24 Dual indexes for MicroPlex Kit v3</a></td>
<td class="format">48 rxns</td>
<td><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010004</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set I</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010005</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set II</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010006</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set III</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010007</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set IV</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</li>
</ul>
</div>
</li>
</ul>
</div>
</div>
</div>
<div class="content active" id="panel2">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<p>The TAG Kit for ChIPmentation offers an optimized ChIP-seq library preparation solution based on tagmentation. This kit includes reagents for tagmentation-based library preparation integrated in the IP and is compatible with any ChIP protocol based on magnetic beads. The primer indexes for multiplexing must be purchased separately and are available as a reference: <a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" target="_blank">24 SI for ChIPmentation</a>, Cat. No. C01011031. Alternatively, for histone marks, Diagenode proposes the complete solution (including all buffers for ChIP, tagmentation and multiplexing): <a href="https://www.diagenode.com/en/p/manual-chipmentation-kit-for-histones-24-rxns" target="_blank">ChIPmentation for Histones</a>.</p>
</div>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> Sample: chromatin-antibody-magnetic beads complexes</li>
<li><i class="fa fa-arrow-circle-right"></i> Input: chromatin from 5 K – 4 M cells</li>
<li><i class="fa fa-arrow-circle-right"></i> Easy and fast protocol</li>
<li><i class="fa fa-arrow-circle-right"></i> Compatible with any ChIP protocol based on magnetic beads</li>
<li><i class="fa fa-arrow-circle-right"></i> No adapter dimers</li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<p class="lead"><em><strong>TAG kit for ChIPmentation</strong></em></p>
<ul class="square">
<li>ChIPmentation library preparation</li>
</ul>
<p class="lead"><em><strong>24 SI for for ChIPmentation</strong></em></p>
<ul class="square">
<li>ChIPmentation library preparation</li>
<li>Tagmentation-based library preparation methods like ATAC-seq, CUT&Tag</li>
</ul>
</div>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C01011030</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" style="color: #b21329;" target="_blank">TAG Kit for ChIPmentation</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C01011031</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" style="color: #b21329;" target="_blank">24 SI for ChIPmentation</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
<div class="content" id="panel3">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<h3 class="text-center diacol"><em>How to choose your library preparation kit?</em></h3>
</div>
<table class="noborder">
<tbody>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Sample</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Chromatin-antibody-beads complex</p>
</td>
<td colspan="2">
<p class="text-center" style="font-size: 15px;">Purified DNA</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Purified DNA</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td colspan="2"><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Application</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">ChIPmentation</p>
</td>
<td colspan="2">
<p class="text-center" style="font-size: 15px;">ChIP-seq library prep<br /> Low input DNA sequencing</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">MeDIP-seq library prep<br /> Genomic DNA sequencing<br /> High input ChIP-seq</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td colspan="2"><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Input</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Chromatin: 5 K to 4 M cells</p>
</td>
<td colspan="2"">
<p class="text-center" style="font-size: 15px;">DNA: 50 pg – 50 ng</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">DNA: 5 ng – 1 µg</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow-45-left.png" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow-45-right.png" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Multiplexing</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 24 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 384 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 48 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 24 samples</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Indexes</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Dual indexes (DI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Kit</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>TAG Kit for ChIPmentation</strong><br /> (indexes not included in the kit)</p>
<p class="text-center"><strong>Kit</strong><br /> <a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" target="_blank">C01011030 – 24 rxns</a></p>
<p class="text-center"><strong>Single indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" target="_blank">C01011031 – 24 SI/24 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>MicroPlex Library Preparation Kit v3</strong><br />(dual indexes not included in the kit)</p>
<p class="text-center"><strong>Kit</strong><br /> <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" target="_blank">C05010001 - 48 rxns</a><br /> <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" target="_blank">C05010002 - 96 rxns</a></p>
<br />
<p class="text-center"><strong>Unique dual indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1" target="_blank">C05010008 - Set I 24 UDI / 24 rxns</a><br /> <a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2" target="_blank">C05010009 - Set II 24 UDI/ 24 rxns</a></p>
<p class="text-center"><strong>Dual indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" target="_blank">C05010003 - 24 DI/ 48 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" target="_blank">C05010004 - Set I 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" target="_blank">C05010005 - Set II 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" target="_blank">C05010006 - Set III 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" target="_blank">C05010007 - Set IV 96 DI/ 96 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>MicroPlex Library Preparation Kit v2</strong><br />(single indexes included in the kit)</p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" target="_blank">C05010012 - 12 SI/ 12 rxns</a><br /> <a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x48-12-indices-48-rxns" target="_blank">C05010013 - 12 SI/ 48 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>iDeal Library Preparation Kit</strong><br />(Set 1 of indexes included in the kit)</p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" target="_blank">C05010020 - 12 SI/ 24 rxns</a></p>
<p class="text-center" style="font-size: 15px;"><strong>Index Primer Set 2</strong></p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" target="_blank">C05010021 - 12 SI/ 24 rxns</a></p>
</td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
<div class="content" id="panel4">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p>Combined chromatin immunoprecipitation and next-generation sequencing (ChIP-seq) has become the gold standard to investigate genome-wide epigenetic profiles. However, ChIP from a limited amount of cells has been a challenge. Here we provide a complete and robust workflow solution for successful ChIP-seq from small numbers of cells using the True MicroChIP kit and MicroPlex Library Preparation kit.</p>
<blockquote><span class="label-green" style="margin-bottom: 16px; margin-left: -22px;">APPLICATION NOTE</span>
<div class="row">
<div class="small-12 medium-6 large-6 columns"><center><img src="https://www.diagenode.com/img/categories/microplex/chip-efficiency-on-10000-cells.jpg" /></center>
<p><small><em>ChIP efficiency on 10,000 cells</em></small></p>
</div>
<div class="small-12 medium-6 large-6 columns">
<p><strong>From minuscule amounts to magnificent results:</strong><br /> reliable ChIP-seq data from 10,000 cells with the True MicroChIP™ and the MicroPlex Library Preparation™ kits.</p>
<a href="https://www.diagenode.com/files/application_notes/True_MicroChIP_and_MicroPlex_kits_Application_Note.pdf" class="details small button" target="_blank">DOWNLOAD</a></div>
</div>
</blockquote>
<blockquote><span class="label-green" style="margin-bottom: 16px; margin-left: -22px;">APPLICATION NOTE</span>
<div class="row">
<div class="small-12 medium-6 large-6 columns"><center><img src="https://www.diagenode.com/img/categories/microplex/quality-control-check.jpg" /></center>
<p class="text-left"><small><em>Quality control check of a ChIP-seq library on the Fragment Analyzer. High Efficiency ChIP performed on 10,000 cells</em></small></p>
</div>
<div class="small-12 medium-6 large-6 columns">
<p class="text-left"><strong>Best Workflow Practices for ChIP-seq Analysis with Small Samples</strong></p>
<a href="https://www.diagenode.com/files/application_notes/Diagenode_AATI_Joint.pdf" class="details small button" target="_blank">DOWNLOAD</a></div>
</div>
</blockquote>
</div>
</div>
</div>
<div class="content" id="panel5">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<h2>TAG Kit for ChIPmentation</h2>
<ol>
<li><strong>What is the difference between tagmentation and ChIPmentation?</strong><br />Tagmentation is a reaction where an enzyme (a transposase) cleaves DNA and incorporates sequencing adaptors at the ends of the fragments in one step. In our ChIPmentation technology we combine chromatin immunoprecipitation and tagmentation in one streamlined workflow where the tagmentation step occurs directly on chromatin.<br /><br /></li>
<li><strong>What is the expected concentration of ChIPmentation libraries?</strong><br />The concentration of libraries that you need to reach will depend on the sensitivity of the machine and kits that you will use to perform the quality control and the sequencing of your libraries. Usually a concentration of 4-8 ng/μl is enough for a quality control using the Qubit High Sensitivity assay (ThermoFischer Scientific) and the High Sensitivity chip for BioAnalyzer (Agilent) and for sequencing on Illumina HiSeq3000/4000.<br /><br /></li>
<li><strong>Does the ChIPmentation approach work on plants?</strong><br />Our ChIPmentation solution has been validated on human cells and we do not have any data on plants. It should be compatible. We would recommend using our Universal Plant ChIP Kit in combination with the TAG Kit for ChIPmentation and the 24 SI for ChIPmentation.<br /><br /></li>
<li><strong>What is the size of the fragments after the tagmentation?</strong><br />The size of the fragments at the end of the ChIPmentation protocol can vary depending on many parameters like the shearing efficiency, the antibody used or the tagmentation time. However, with our standard protocol we usually obtain a library peak which is around 200-300 bp (see example of results at the end of the manual). If many fragments larger than 500 bp are present , the best would be to contact your sequencing provider to ask what their requirements are, because it can vary depending on the sequencer. If you want to remove the large fragments you can use the size selection protocol described in the manual.<br /><br /></li>
<li><strong>What is the size of the adapters?</strong><br />The sum of the adapters is 128 bp.</li>
</ol>
</div>
<div class="extra-spaced">
<h2>MicroPlex Library Preparation Kit</h2>
<ol>
<li><strong>Can I use the available Illumina primers and validate them with the MicroPlex Kit v2?</strong><br /> Although the final flanking sequences of MicroPlex are the same as those used by Illumina, the PCR primers are not identical and part of them is supplied with the buffer. For this reason Illumina primers will not work as substitute.<br /><br /></li>
<li><strong>The BioAnalyzer profile of purified library shows the presence of low molecular weight peaks (primers/adaptors) in the samples. Should I re- purify the samples or they can be used directly to the sequencing? If the second purification is recommended, which ratio sample/AMPure beads should I use?</strong><br /> You can do a second round of purification using 1:1 ratio of AMPure beads to sample and this should get rid of the majority of the dimers.<br /><br /></li>
<li><strong>I am going to use the MicroPlex Library Preparation Kit v2 on ChIP samples . Our thermocycler has ramp rate 1.5°/s max while the protocol recommends using a ramp rate 3 to 5°/s. How would this affect the library prep?</strong><br /> We have not used a thermocycler with a ramp rate of 1.5 °C, which seems faster than most of thermocyclers. Too fast of a ramp rate may affect the primer annealing and ligation steps.<br /><br /></li>
<li><strong>What is the function of the replication stop site in the adapter loops?</strong><br /> The replication stop site in the adaptor loops function to stop the polymerase from continuing to copy the rest of the stem loop.<br /><br /></li>
<li><strong>I want to do ChIP-seq. Which ChIP-seq kit can I use for sample preparation prior to Microplex Library Preparation Kit v2?</strong><br /> In our portfolio there are several ChIP-seq kits compatible with Microplex Library Preparation Kit v2. Depending on your sample type and target studied you can use the following kits: iDeal ChIP-seq Kit for Transcription Factors (Cat. No. C01010055), iDeal ChIP-seq Kit for Histones (Cat. No. C01010051), True MicroChIP kit (Cat. No. C01010130), Universal Plant ChIP-seq Kit (Cat. No. C01010152). All these kits exist in manual and automated versions.<br /><br /></li>
<li><strong>Is Microplex Library Preparation Kit v2 compatible with exome enrichment methods?</strong><br /> Microplex Library Preparation Kit v2 is compatible with major exome and target enrichment products, including Agilent SureSelect<sup>®</sup>, Roche NimbleGen<sup>®</sup> SeqCap<sup>®</sup> EZ and custom panels.<br /><br /></li>
<li><strong>What is the nick that is mentioned in the kit method overview?</strong><br /> The nick is simply a gap between a stem adaptor and 3’ DNA end, as shown on the schema in the kit method overview.<br /><br /></li>
<li><strong>Are the indexes of the MicroPlex library preparation kit v2 located at i5 or i7?</strong><br /> The libraries generated with the MicroPlex kit v2 contain indices located at i7.<br /><br /></li>
<li><strong>Is there a need to use custom index read primers for the sequencing to read the 8nt iPCRtags?</strong><br /> There is no need for using custom Sequencing primer to sequence MicroPlex libraires. MicroPlex libraries can be sequenced using standard Illumina Sequencing kits and protocols.<br /><br /></li>
<li><strong>What is the advantage of using stem-loop adapter in the MicroPlex kit?</strong><br /> There are several advantages of using stem-loop adaptors. First of all, stem-loop adaptors prevent from self-ligation thus increases the ligation efficiency between the adapter and DNA fragment. Moreover, the background is reduced using ds adaptors with no single-stranded tails. Finally, adaptor-adaptor ligation is reduced using blocked 5’ ends.<br /><br /></li>
</ol>
</div>
<div class="extra-spaced">
<h2>IDeal Library Preparation Kit</h2>
<ol>
<li><strong>Are the index from the iDeal library Prep kit compatible with the MicroPlex library prep kit?</strong><br /> No, it is important to use only the indexes provided in the MicroPlex kit to ensure proper library preparation with this kit</li>
</ol>
</div>
</div>
</div>
</div>
</div>
</div>
</div>
<script>// <![CDATA[
$(document).ready(function() {
$('ul.tips-menu li a:first').trigger('focus');
});
// ]]></script>',
'no_promo' => false,
'in_menu' => true,
'online' => true,
'tabular' => true,
'hide' => true,
'all_format' => false,
'is_antibody' => false,
'slug' => 'library-preparation-for-ChIP-seq',
'cookies_tag_id' => null,
'meta_keywords' => 'Library preparation,Next Generation Sequencing,DNA fragments,MicroPlex Library,iDeal Library',
'meta_description' => 'Diagenode Kits have been Optimized for DNA library Preparation used for Next Generation Sequencing for a range of inputs.',
'meta_title' => 'DNA Library Preparation Kits,Microplex library Preparation Kits | Diagenode',
'modified' => '2021-12-20 14:30:28',
'created' => '2016-10-21 02:39:19',
'ProductsCategory' => array(
[maximum depth reached]
),
'CookiesTag' => array([maximum depth reached])
)
),
'Document' => array(
(int) 0 => array(
'id' => '1056',
'name' => 'Primer indexes for MicroPlex kit v3',
'description' => '<p>High Performance Library Preparation for Illumina<sup>®</sup> NGS Platforms</p>',
'image_id' => null,
'type' => 'Manual',
'url' => 'files/products/kits/Primer-indexes-for-Microplex-v3.pdf',
'slug' => 'primer-indexes-for-microplex-v3-manual',
'meta_keywords' => '',
'meta_description' => '',
'modified' => '2020-10-22 12:03:45',
'created' => '2019-07-18 11:53:57',
'ProductsDocument' => array(
[maximum depth reached]
)
)
),
'Feature' => array(),
'Image' => array(
(int) 0 => array(
'id' => '1759',
'name' => 'product/kits/chip-kit-icon.png',
'alt' => 'chip kit icon',
'modified' => '2018-01-18 15:26:13',
'created' => '2018-01-18 15:08:38',
'ProductsImage' => array(
[maximum depth reached]
)
)
),
'Promotion' => array(),
'Protocol' => array(),
'Publication' => array(),
'Testimonial' => array(),
'Area' => array(),
'SafetySheet' => array()
)
$country = 'US'
$countries_allowed = array(
(int) 0 => 'CA',
(int) 1 => 'US',
(int) 2 => 'IE',
(int) 3 => 'GB',
(int) 4 => 'DK',
(int) 5 => 'NO',
(int) 6 => 'SE',
(int) 7 => 'FI',
(int) 8 => 'NL',
(int) 9 => 'BE',
(int) 10 => 'LU',
(int) 11 => 'FR',
(int) 12 => 'DE',
(int) 13 => 'CH',
(int) 14 => 'AT',
(int) 15 => 'ES',
(int) 16 => 'IT',
(int) 17 => 'PT'
)
$outsource = false
$other_formats = array()
$edit = ''
$testimonials = ''
$featured_testimonials = ''
$related_products = '<li>
<div class="row">
<div class="small-12 columns">
<a href="/en/p/microplex-lib-prep-kit-v3-48-rxns"><img src="/img/product/kits/chip-kit-icon.png" alt="chip kit icon" class="th"/></a> </div>
<div class="small-12 columns">
<div class="small-6 columns" style="padding-left:0px;padding-right:0px;margin-top:-6px;margin-left:-1px">
<span class="success label" style="">C05010001</span>
</div>
<div class="small-6 columns text-right" style="padding-left:0px;padding-right:0px;margin-top:-6px">
<!--a href="#" style="color:#B21329"><i class="fa fa-info-circle"></i></a-->
<!-- BEGIN: ADD TO CART MODAL --><div id="cartModal-3032" class="reveal-modal small" data-reveal aria-labelledby="modalTitle" aria-hidden="true" role="dialog">
<form action="/en/carts/add/3032" id="CartAdd/3032Form" method="post" accept-charset="utf-8"><div style="display:none;"><input type="hidden" name="_method" value="POST"/></div><input type="hidden" name="data[Cart][product_id]" value="3032" id="CartProductId"/>
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p>Add <input name="data[Cart][quantity]" placeholder="1" value="1" min="1" style="width:60px;display:inline" type="number" id="CartQuantity" required="required"/> <strong> MicroPlex Library Preparation Kit v3 /48 rxns</strong> to my shopping cart.</p>
<div class="row">
<div class="small-6 medium-6 large-6 columns">
<button class="alert small button expand" onclick="$(this).addToCart('MicroPlex Library Preparation Kit v3 /48 rxns',
'C05010001',
'1555',
$('#CartQuantity').val());" name="checkout" id="checkout" value="checkout" type="submit">Checkout</button> </div>
<div class="small-6 medium-6 large-6 columns">
<button class="alert small button expand" onclick="$(this).addToCart('MicroPlex Library Preparation Kit v3 /48 rxns',
'C05010001',
'1555',
$('#CartQuantity').val());" name="keepshop" id="keepshop" type="submit">Keep shopping</button> </div>
</div>
</div>
</div>
</form><a class="close-reveal-modal" aria-label="Close">×</a></div><!-- END: ADD TO CART MODAL --><a href="#" id="microplex-lib-prep-kit-v3-48-rxns" data-reveal-id="cartModal-3032" class="" style="color:#B21329"><i class="fa fa-cart-plus"></i></a>
</div>
</div>
<div class="small-12 columns" >
<h6 style="height:60px">MicroPlex Library Preparation Kit v3 /48 rxns</h6>
</div>
</div>
</li>
'
$related = array(
'id' => '3032',
'antibody_id' => null,
'name' => 'MicroPlex Library Preparation Kit v3 /48 rxns',
'description' => '<p><a href="https://www.diagenode.com/files/products/kits/Microplex-library-prep-v3.pdf"><img src="https://www.diagenode.com/img/buttons/bt-manual.png" /></a></p>
<p>Diagenode’s <strong>MicroPlex Library Preparation Kits v3</strong> have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg. The kit MicroPlex v3 includes all buffers and enzymes necessary for the library preparation. For flexibility of the choice different formats of compatible primer indexes are available separately:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2">C05010005 - 96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p style="padding-left: 30px;">NEW! Unique dual indexes :</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1">C05010008 - 24 UDI for MicroPlex Kit v3 - Set I</a></li>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2">C05010009 - 24 UDI for MicroPlex Kit v3 - Set II</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong> with 24 dual indexes (8 nt)</li>
<li><strong>Validated with the IP-<a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">Star<sup>®</sup></a></strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit"> Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img alt="MicroPlex Library Preparation Kit v3 /48 rxns" src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label2' => '',
'info2' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label3' => '',
'info3' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'format' => '48 rxns',
'catalog_number' => 'C05010001',
'old_catalog_number' => '',
'sf_code' => 'C05010001-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1750',
'price_USD' => '1555',
'price_GBP' => '1555',
'price_JPY' => '286740',
'price_CNY' => '',
'price_AUD' => '3888',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => 'microplex-lib-prep-kit-v3-48-rxns',
'meta_title' => 'MicroPlex Library Preparation Kit v3 /48 rxns | Diagenode',
'meta_keywords' => 'MicroPlex Library Preparation Kit',
'meta_description' => 'MicroPlex Library Preparation Kits v3 have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg.',
'modified' => '2023-04-06 12:08:35',
'created' => '2019-07-16 16:03:58',
'ProductsRelated' => array(
'id' => '4541',
'product_id' => '3027',
'related_id' => '3032'
),
'Image' => array(
(int) 0 => array(
'id' => '1759',
'name' => 'product/kits/chip-kit-icon.png',
'alt' => 'chip kit icon',
'modified' => '2018-01-18 15:26:13',
'created' => '2018-01-18 15:08:38',
'ProductsImage' => array(
[maximum depth reached]
)
)
)
)
$rrbs_service = array(
(int) 0 => (int) 1894,
(int) 1 => (int) 1895
)
$chipseq_service = array(
(int) 0 => (int) 2683,
(int) 1 => (int) 1835,
(int) 2 => (int) 1836,
(int) 3 => (int) 2684,
(int) 4 => (int) 1838,
(int) 5 => (int) 1839,
(int) 6 => (int) 1856
)
$labelize = object(Closure) {
}
$old_catalog_number = ''
$country_code = 'US'
$label = '<img src="/img/banners/banner-customizer-back.png" alt=""/>'
$document = array(
'id' => '1056',
'name' => 'Primer indexes for MicroPlex kit v3',
'description' => '<p>High Performance Library Preparation for Illumina<sup>®</sup> NGS Platforms</p>',
'image_id' => null,
'type' => 'Manual',
'url' => 'files/products/kits/Primer-indexes-for-Microplex-v3.pdf',
'slug' => 'primer-indexes-for-microplex-v3-manual',
'meta_keywords' => '',
'meta_description' => '',
'modified' => '2020-10-22 12:03:45',
'created' => '2019-07-18 11:53:57',
'ProductsDocument' => array(
'id' => '2775',
'product_id' => '3027',
'document_id' => '1056'
)
)include - APP/View/Products/view.ctp, line 755
View::_evaluate() - CORE/Cake/View/View.php, line 971
View::_render() - CORE/Cake/View/View.php, line 933
View::render() - CORE/Cake/View/View.php, line 473
Controller::render() - CORE/Cake/Controller/Controller.php, line 963
ProductsController::slug() - APP/Controller/ProductsController.php, line 1052
ReflectionMethod::invokeArgs() - [internal], line ??
Controller::invokeAction() - CORE/Cake/Controller/Controller.php, line 491
Dispatcher::_invoke() - CORE/Cake/Routing/Dispatcher.php, line 193
Dispatcher::dispatch() - CORE/Cake/Routing/Dispatcher.php, line 167
[main] - APP/webroot/index.php, line 118
Notice (8): Undefined variable: header [APP/View/Products/view.ctp, line 755]Code Context<!-- BEGIN: REQUEST_FORM MODAL -->
<div id="request_formModal" class="reveal-modal medium" data-reveal aria-labelledby="modalTitle" aria-hidden="true" role="dialog">
<?= $this->element('Forms/simple_form', array('solution_of_interest' => $solution_of_interest, 'header' => $header, 'message' => $message, 'campaign_id' => $campaign_id)) ?>
$viewFile = '/home/website-server/www/app/View/Products/view.ctp'
$dataForView = array(
'language' => 'en',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'product' => array(
'Product' => array(
'id' => '3027',
'antibody_id' => null,
'name' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'description' => '<p>Diagenode’s <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns">MicroPlex Library Preparation Kits v3</a> have been extensively validated for ChIP-seq library prep from ChIP-derived DNA. The kit MicroPlex v3 has to be purchased with the compatible set of dual indexes. This set of dual indexes allows for multiplexing up to 96 samples.</p>
<p>Check all available sets of dual indexes:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong></li>
<li><strong>Validated with the </strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">IP-Star Compact Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>',
'label2' => '',
'info2' => '',
'label3' => '',
'info3' => '',
'format' => '96 rxns',
'catalog_number' => 'C05010005',
'old_catalog_number' => '',
'sf_code' => 'C05010005-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1355',
'price_USD' => '1300',
'price_GBP' => '1235',
'price_JPY' => '222015',
'price_CNY' => '',
'price_AUD' => '3250',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => '96-dual-indexes-for-microplex-kit-v3-set-2',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'modified' => '2020-10-23 17:11:41',
'created' => '2019-07-16 15:48:28',
'locale' => 'eng'
),
'Antibody' => array(
'host' => '*****',
'id' => null,
'name' => null,
'description' => null,
'clonality' => null,
'isotype' => null,
'lot' => null,
'concentration' => null,
'reactivity' => null,
'type' => null,
'purity' => null,
'classification' => null,
'application_table' => null,
'storage_conditions' => null,
'storage_buffer' => null,
'precautions' => null,
'uniprot_acc' => null,
'slug' => null,
'meta_keywords' => null,
'meta_description' => null,
'modified' => null,
'created' => null,
'select_label' => null
),
'Slave' => array(),
'Group' => array(),
'Related' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Application' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Category' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Document' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Feature' => array(),
'Image' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Promotion' => array(),
'Protocol' => array(),
'Publication' => array(),
'Testimonial' => array(),
'Area' => array(),
'SafetySheet' => array()
)
)
$language = 'en'
$meta_keywords = ''
$meta_description = '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns'
$meta_title = '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns'
$product = array(
'Product' => array(
'id' => '3027',
'antibody_id' => null,
'name' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'description' => '<p>Diagenode’s <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns">MicroPlex Library Preparation Kits v3</a> have been extensively validated for ChIP-seq library prep from ChIP-derived DNA. The kit MicroPlex v3 has to be purchased with the compatible set of dual indexes. This set of dual indexes allows for multiplexing up to 96 samples.</p>
<p>Check all available sets of dual indexes:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong></li>
<li><strong>Validated with the </strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">IP-Star Compact Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>',
'label2' => '',
'info2' => '',
'label3' => '',
'info3' => '',
'format' => '96 rxns',
'catalog_number' => 'C05010005',
'old_catalog_number' => '',
'sf_code' => 'C05010005-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1355',
'price_USD' => '1300',
'price_GBP' => '1235',
'price_JPY' => '222015',
'price_CNY' => '',
'price_AUD' => '3250',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => '96-dual-indexes-for-microplex-kit-v3-set-2',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'modified' => '2020-10-23 17:11:41',
'created' => '2019-07-16 15:48:28',
'locale' => 'eng'
),
'Antibody' => array(
'host' => '*****',
'id' => null,
'name' => null,
'description' => null,
'clonality' => null,
'isotype' => null,
'lot' => null,
'concentration' => null,
'reactivity' => null,
'type' => null,
'purity' => null,
'classification' => null,
'application_table' => null,
'storage_conditions' => null,
'storage_buffer' => null,
'precautions' => null,
'uniprot_acc' => null,
'slug' => null,
'meta_keywords' => null,
'meta_description' => null,
'modified' => null,
'created' => null,
'select_label' => null
),
'Slave' => array(),
'Group' => array(),
'Related' => array(
(int) 0 => array(
'id' => '3032',
'antibody_id' => null,
'name' => 'MicroPlex Library Preparation Kit v3 /48 rxns',
'description' => '<p><a href="https://www.diagenode.com/files/products/kits/Microplex-library-prep-v3.pdf"><img src="https://www.diagenode.com/img/buttons/bt-manual.png" /></a></p>
<p>Diagenode’s <strong>MicroPlex Library Preparation Kits v3</strong> have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg. The kit MicroPlex v3 includes all buffers and enzymes necessary for the library preparation. For flexibility of the choice different formats of compatible primer indexes are available separately:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2">C05010005 - 96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p style="padding-left: 30px;">NEW! Unique dual indexes :</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1">C05010008 - 24 UDI for MicroPlex Kit v3 - Set I</a></li>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2">C05010009 - 24 UDI for MicroPlex Kit v3 - Set II</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong> with 24 dual indexes (8 nt)</li>
<li><strong>Validated with the IP-<a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">Star<sup>®</sup></a></strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit"> Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img alt="MicroPlex Library Preparation Kit v3 /48 rxns" src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label2' => '',
'info2' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label3' => '',
'info3' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'format' => '48 rxns',
'catalog_number' => 'C05010001',
'old_catalog_number' => '',
'sf_code' => 'C05010001-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1750',
'price_USD' => '1555',
'price_GBP' => '1555',
'price_JPY' => '286740',
'price_CNY' => '',
'price_AUD' => '3888',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => 'microplex-lib-prep-kit-v3-48-rxns',
'meta_title' => 'MicroPlex Library Preparation Kit v3 /48 rxns | Diagenode',
'meta_keywords' => 'MicroPlex Library Preparation Kit',
'meta_description' => 'MicroPlex Library Preparation Kits v3 have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg.',
'modified' => '2023-04-06 12:08:35',
'created' => '2019-07-16 16:03:58',
'ProductsRelated' => array(
[maximum depth reached]
),
'Image' => array(
[maximum depth reached]
)
)
),
'Application' => array(
(int) 0 => array(
'id' => '14',
'position' => '10',
'parent_id' => '3',
'name' => 'DNA/RNA library preparation',
'description' => '<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p><span style="font-weight: 400;">Most of the major next-generation sequencing platforms require ligation of specific adaptor oligos to </span><a href="../applications/dna-rna-shearing"><span style="font-weight: 400;">fragmented DNA or RNA</span></a><span style="font-weight: 400;"> prior to sequencing</span></p>
<p><span style="font-weight: 400;">After input DNA has been fragmented, it is end-repaired and blunt-ended</span><span style="font-weight: 400;">. The next step is a A-tailing in which dAMP is added to the 3´ end of the blunt phosphorylated DNA fragments to prevent concatemerization and to allow the ligation of adaptors with complementary dT overhangs. In addition, barcoded adapters can be incorporated to facilitate multiplexing prior to or during amplification.</span></p>
<center><img src="https://www.diagenode.com/img/categories/library-prep/flux.png" /></center>
<p><span style="font-weight: 400;">Diagenode offers a comprehensive product portfolio for library preparation:<br /></span></p>
<strong><a href="https://www.diagenode.com/en/categories/Library-preparation-for-RNA-seq">D-Plex RNA-seq Library Preparation Kits</a></strong><br />
<p><span style="font-weight: 400;">Diagenode’s new RNA-sequencing solutions utilize the innovative c</span><span style="font-weight: 400;">apture and a</span><span style="font-weight: 400;">mplification by t</span><span style="font-weight: 400;">ailing and s</span><span style="font-weight: 400;">witching”</span><span style="font-weight: 400;">, a ligation-free method to produce DNA libraries for next generation sequencing from low input amounts of RNA. </span><span style="font-weight: 400;"></span><a href="../categories/Library-preparation-for-RNA-seq">Learn more</a></p>
<strong><a href="../categories/library-preparation-for-ChIP-seq">ChIP-seq and DNA sequencing library preparation solutions</a></strong><br />
<p><span style="font-weight: 400;">Our kits have been optimized for DNA library preparation used for next generation sequencing for a wide range of inputs. Using a simple three-step protocols, our</span><a href="http://www.diagenode.com/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns"><span style="font-weight: 400;"> </span></a><span style="font-weight: 400;">kits are an optimal choice for library preparation from DNA inputs down to 50 pg. </span><a href="../categories/library-preparation-for-ChIP-seq">Learn more</a></p>
<a href="../p/bioruptor-pico-sonication-device"><span style="font-weight: 400;"></span><strong>Bioruptor Pico - short fragments</strong></a><a href="../categories/library-preparation-for-ChIP-seq-and-DNA-sequencing"><span style="font-weight: 400;"></span></a><br />
<p><span style="font-weight: 400;"></span><span style="font-weight: 400;">Our well-cited Bioruptor Pico is the shearing device of choice for chromatin and DNA fragmentation. Obtain uniform and tight fragment distributions between 150bp -2kb. </span><a href="../p/bioruptor-pico-sonication-device">Learn more</a></p>
<strong><a href="../p/megaruptor2-1-unit"><span href="../p/bioruptor-pico-sonication-device">Megaruptor</span>® - long fragments</a></strong><a href="../p/bioruptor-pico-sonication-device"><span style="font-weight: 400;"></span></a><a href="../categories/library-preparation-for-ChIP-seq-and-DNA-sequencing"><span style="font-weight: 400;"></span></a><br />
<p><span style="font-weight: 400;"></span><span style="font-weight: 400;">The Megaruptor is designed to shear DNA from 3kb-75kb for long-read sequencing. <a href="../p/megaruptor2-1-unit">Learn more</a></span></p>
<span href="../p/bioruptor-pico-sonication-device"></span><span style="font-weight: 400;"></span></div>
</div>',
'in_footer' => false,
'in_menu' => true,
'online' => true,
'tabular' => true,
'slug' => 'library-preparation',
'meta_keywords' => 'Library preparation,Next Generation Sequencing,DNA fragments,MicroPlex Library,iDeal Library',
'meta_description' => 'Diagenode offers a comprehensive product portfolio for library preparation such as CATS RNA-seq Library Preparation Kits,ChIP-seq and DNA sequencing library preparation solutions',
'meta_title' => 'Library preparation for Next Generation Sequencing (NGS) | Diagenode',
'modified' => '2021-03-10 03:27:16',
'created' => '2014-08-16 07:47:56',
'ProductsApplication' => array(
[maximum depth reached]
)
)
),
'Category' => array(
(int) 0 => array(
'id' => '124',
'position' => '1',
'parent_id' => '15',
'name' => 'Library preparation for ChIP-seq',
'description' => '<div class="row">
<div class="large-12 columns text-justify">
<p>Library preparation following ChIP can be challenging due to the limited amount of DNA recovered after ChIP. Diagenode has developed the optimal solutions for ChIP-seq using two different approaches: the ligation-based library preparation on purified DNA or the tagmentation-based ChIPmentation.</p>
</div>
</div>
<div class="row extra-spaced">
<div class="large-12 columns"><center><a href="https://www.diagenode.com/en/pages/form-microplex-promo" target="_blank"></a></center></div>
</div>
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div id="portal" class="main-portal">
<div class="portal-inner"><nav class="portal-nav">
<ul data-tab="" class="tips-menu">
<li><a href="#panel1" class="tips portal button">Ligation-based library prep</a></li>
<li><a href="#panel2" class="tips portal button">ChIPmentation</a></li>
<li><a href="#panel3" class="tips portal button">Kit choice guide</a></li>
<li><a href="#panel4" class="tips portal button">Resources</a></li>
<li><a href="#panel5" class="tips portal button">FAQs</a></li>
</ul>
</nav></div>
</div>
<div class="tabs-content">
<div class="content active" id="panel1">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v5" style="color: #13b29c;"><i class="fa fa-caret-right"></i> Standard input library prep</a>
<div id="v5" class="content">
<div class="small-12 medium-12 large-12 columns">
<p>The <strong>iDeal Library Preparation Kit</strong> reliably converts DNA into indexed libraries for next-generation sequencing, with input amounts down to <strong>5 ng</strong>. Our kit offers a simple and fast workflow, high yields, and ready-to-sequence DNA on the Illumina platform.</p>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> <strong>Sample</strong>: Fragmented dsDNA</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Input</strong>: 5 ng – 1 µg</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Fast protocol</strong>: 3 hours</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Easy processing</strong>: 3 steps</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Indexing</strong>: single indexes for multiplexing up to 24 samples</li>
<li><i class="fa fa-arrow-circle-right"></i> Manual and automated protocols available</li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<ul class="square">
<li>MeDIP-seq library prep</li>
<li>Genomic DNA sequencing</li>
<li>High input ChIP-seq</li>
</ul>
</div>
<div class="extra-spaced">
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010020</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" style="color: #b21329;" target="_blank">iDeal Library Preparation Kit x24 (incl. Index Primer Set 1)</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010021</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" style="color: #b21329;" target="_blank">Index Primer Set 2 (iDeal Lib. Prep Kit x24)</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
</li>
</ul>
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v4" style="color: #13b29c;"><i class="fa fa-caret-right"></i> Low input library prep</a>
<div id="v4" class="content active"><center><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" target="_blank"><img src="https://www.diagenode.com/img/banners/banner-microplex-v3-580.jpg" class="extra-spaced" /></a></center>
<div align="center"><a href="https://www.diagenode.com/pages/form-microplex3" class="center alert radius button extra-spaced"><i class="fa fa-info"></i> Contact us</a></div>
<div class="extra-spaced">
<p>Diagenode’s <strong>MicroPlex Library Preparation kits</strong> have been extensively validated for ChIP-seq samples. Generated libraries are compatible with single-end or paired-end sequencing. MicroPlex chemistry (using stem-loop adapters ) is specifically developed and optimized to generate DNA libraries with high molecular complexity from the lowest input amounts. Only <strong>50 pg to 50 ng</strong> of fragmented double-stranded DNA is required for library preparation. The entire <strong>three-step workflow</strong> takes place in a <strong>single tube</strong> or well in about <strong>2 hours</strong>. No intermediate purification steps and no sample transfers are necessary to prevent handling errors and loss of valuable samples.</p>
</div>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> <strong>Sample</strong>: Fragmented dsDNA</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Low input</strong>: 50 pg – 50 ng</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Fast protocol</strong>: 2 hours</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Easy processing</strong>: 3 steps in 1 tube</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>No intermediate purification</strong></li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
<li><i class="fa fa-arrow-circle-right"></i> Manual and automated protocols available</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<ul class="square">
<li>ChIP-seq library prep from ChIP-derived DNA</li>
<li>Low input DNA sequencing</li>
</ul>
</div>
<h2>Two versions are available:</h2>
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v2" style="color: #13b29c;"><i class="fa fa-caret-right"></i> MicroPlex Library Preparation Kit v2 with single indexes</a>
<div id="v2" class="content">
<p>The MicroPlex Library Preparation Kit v2 contains all necessary reagents including single indexes for multiplexing up to 48 samples using single barcoding.</p>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010012</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v2 (12 indexes)</a></td>
<td class="format">12 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</li>
<li class="accordion-navigation"><a href="#v3" style="color: #13b29c;"><i class="fa fa-caret-right"></i> MicroPlex Library Preparation Kit v3 with dual indexes <strong><span class="diacol">NEW!</span></strong></a>
<div id="v3" class="content active">
<p>In this version the library preparation reagents and the dual indexes are available separately allowing for the flexibility choosing the number of indexes. MicroPlex v3 has multiplexing capacities up to 384 samples.</p>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010001</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v3 /48 rxns</a></td>
<td class="format">48 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010002</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v3 /96 rxns</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
<h4>DUAL INDEXES</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010003</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" style="color: #b21329;" target="_blank">24 Dual indexes for MicroPlex Kit v3</a></td>
<td class="format">48 rxns</td>
<td><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010004</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set I</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010005</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set II</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010006</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set III</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010007</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set IV</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</li>
</ul>
</div>
</li>
</ul>
</div>
</div>
</div>
<div class="content active" id="panel2">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<p>The TAG Kit for ChIPmentation offers an optimized ChIP-seq library preparation solution based on tagmentation. This kit includes reagents for tagmentation-based library preparation integrated in the IP and is compatible with any ChIP protocol based on magnetic beads. The primer indexes for multiplexing must be purchased separately and are available as a reference: <a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" target="_blank">24 SI for ChIPmentation</a>, Cat. No. C01011031. Alternatively, for histone marks, Diagenode proposes the complete solution (including all buffers for ChIP, tagmentation and multiplexing): <a href="https://www.diagenode.com/en/p/manual-chipmentation-kit-for-histones-24-rxns" target="_blank">ChIPmentation for Histones</a>.</p>
</div>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> Sample: chromatin-antibody-magnetic beads complexes</li>
<li><i class="fa fa-arrow-circle-right"></i> Input: chromatin from 5 K – 4 M cells</li>
<li><i class="fa fa-arrow-circle-right"></i> Easy and fast protocol</li>
<li><i class="fa fa-arrow-circle-right"></i> Compatible with any ChIP protocol based on magnetic beads</li>
<li><i class="fa fa-arrow-circle-right"></i> No adapter dimers</li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<p class="lead"><em><strong>TAG kit for ChIPmentation</strong></em></p>
<ul class="square">
<li>ChIPmentation library preparation</li>
</ul>
<p class="lead"><em><strong>24 SI for for ChIPmentation</strong></em></p>
<ul class="square">
<li>ChIPmentation library preparation</li>
<li>Tagmentation-based library preparation methods like ATAC-seq, CUT&Tag</li>
</ul>
</div>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C01011030</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" style="color: #b21329;" target="_blank">TAG Kit for ChIPmentation</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C01011031</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" style="color: #b21329;" target="_blank">24 SI for ChIPmentation</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
<div class="content" id="panel3">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<h3 class="text-center diacol"><em>How to choose your library preparation kit?</em></h3>
</div>
<table class="noborder">
<tbody>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Sample</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Chromatin-antibody-beads complex</p>
</td>
<td colspan="2">
<p class="text-center" style="font-size: 15px;">Purified DNA</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Purified DNA</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td colspan="2"><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Application</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">ChIPmentation</p>
</td>
<td colspan="2">
<p class="text-center" style="font-size: 15px;">ChIP-seq library prep<br /> Low input DNA sequencing</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">MeDIP-seq library prep<br /> Genomic DNA sequencing<br /> High input ChIP-seq</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td colspan="2"><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Input</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Chromatin: 5 K to 4 M cells</p>
</td>
<td colspan="2"">
<p class="text-center" style="font-size: 15px;">DNA: 50 pg – 50 ng</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">DNA: 5 ng – 1 µg</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow-45-left.png" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow-45-right.png" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Multiplexing</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 24 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 384 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 48 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 24 samples</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Indexes</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Dual indexes (DI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Kit</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>TAG Kit for ChIPmentation</strong><br /> (indexes not included in the kit)</p>
<p class="text-center"><strong>Kit</strong><br /> <a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" target="_blank">C01011030 – 24 rxns</a></p>
<p class="text-center"><strong>Single indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" target="_blank">C01011031 – 24 SI/24 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>MicroPlex Library Preparation Kit v3</strong><br />(dual indexes not included in the kit)</p>
<p class="text-center"><strong>Kit</strong><br /> <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" target="_blank">C05010001 - 48 rxns</a><br /> <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" target="_blank">C05010002 - 96 rxns</a></p>
<br />
<p class="text-center"><strong>Unique dual indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1" target="_blank">C05010008 - Set I 24 UDI / 24 rxns</a><br /> <a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2" target="_blank">C05010009 - Set II 24 UDI/ 24 rxns</a></p>
<p class="text-center"><strong>Dual indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" target="_blank">C05010003 - 24 DI/ 48 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" target="_blank">C05010004 - Set I 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" target="_blank">C05010005 - Set II 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" target="_blank">C05010006 - Set III 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" target="_blank">C05010007 - Set IV 96 DI/ 96 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>MicroPlex Library Preparation Kit v2</strong><br />(single indexes included in the kit)</p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" target="_blank">C05010012 - 12 SI/ 12 rxns</a><br /> <a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x48-12-indices-48-rxns" target="_blank">C05010013 - 12 SI/ 48 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>iDeal Library Preparation Kit</strong><br />(Set 1 of indexes included in the kit)</p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" target="_blank">C05010020 - 12 SI/ 24 rxns</a></p>
<p class="text-center" style="font-size: 15px;"><strong>Index Primer Set 2</strong></p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" target="_blank">C05010021 - 12 SI/ 24 rxns</a></p>
</td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
<div class="content" id="panel4">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p>Combined chromatin immunoprecipitation and next-generation sequencing (ChIP-seq) has become the gold standard to investigate genome-wide epigenetic profiles. However, ChIP from a limited amount of cells has been a challenge. Here we provide a complete and robust workflow solution for successful ChIP-seq from small numbers of cells using the True MicroChIP kit and MicroPlex Library Preparation kit.</p>
<blockquote><span class="label-green" style="margin-bottom: 16px; margin-left: -22px;">APPLICATION NOTE</span>
<div class="row">
<div class="small-12 medium-6 large-6 columns"><center><img src="https://www.diagenode.com/img/categories/microplex/chip-efficiency-on-10000-cells.jpg" /></center>
<p><small><em>ChIP efficiency on 10,000 cells</em></small></p>
</div>
<div class="small-12 medium-6 large-6 columns">
<p><strong>From minuscule amounts to magnificent results:</strong><br /> reliable ChIP-seq data from 10,000 cells with the True MicroChIP™ and the MicroPlex Library Preparation™ kits.</p>
<a href="https://www.diagenode.com/files/application_notes/True_MicroChIP_and_MicroPlex_kits_Application_Note.pdf" class="details small button" target="_blank">DOWNLOAD</a></div>
</div>
</blockquote>
<blockquote><span class="label-green" style="margin-bottom: 16px; margin-left: -22px;">APPLICATION NOTE</span>
<div class="row">
<div class="small-12 medium-6 large-6 columns"><center><img src="https://www.diagenode.com/img/categories/microplex/quality-control-check.jpg" /></center>
<p class="text-left"><small><em>Quality control check of a ChIP-seq library on the Fragment Analyzer. High Efficiency ChIP performed on 10,000 cells</em></small></p>
</div>
<div class="small-12 medium-6 large-6 columns">
<p class="text-left"><strong>Best Workflow Practices for ChIP-seq Analysis with Small Samples</strong></p>
<a href="https://www.diagenode.com/files/application_notes/Diagenode_AATI_Joint.pdf" class="details small button" target="_blank">DOWNLOAD</a></div>
</div>
</blockquote>
</div>
</div>
</div>
<div class="content" id="panel5">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<h2>TAG Kit for ChIPmentation</h2>
<ol>
<li><strong>What is the difference between tagmentation and ChIPmentation?</strong><br />Tagmentation is a reaction where an enzyme (a transposase) cleaves DNA and incorporates sequencing adaptors at the ends of the fragments in one step. In our ChIPmentation technology we combine chromatin immunoprecipitation and tagmentation in one streamlined workflow where the tagmentation step occurs directly on chromatin.<br /><br /></li>
<li><strong>What is the expected concentration of ChIPmentation libraries?</strong><br />The concentration of libraries that you need to reach will depend on the sensitivity of the machine and kits that you will use to perform the quality control and the sequencing of your libraries. Usually a concentration of 4-8 ng/μl is enough for a quality control using the Qubit High Sensitivity assay (ThermoFischer Scientific) and the High Sensitivity chip for BioAnalyzer (Agilent) and for sequencing on Illumina HiSeq3000/4000.<br /><br /></li>
<li><strong>Does the ChIPmentation approach work on plants?</strong><br />Our ChIPmentation solution has been validated on human cells and we do not have any data on plants. It should be compatible. We would recommend using our Universal Plant ChIP Kit in combination with the TAG Kit for ChIPmentation and the 24 SI for ChIPmentation.<br /><br /></li>
<li><strong>What is the size of the fragments after the tagmentation?</strong><br />The size of the fragments at the end of the ChIPmentation protocol can vary depending on many parameters like the shearing efficiency, the antibody used or the tagmentation time. However, with our standard protocol we usually obtain a library peak which is around 200-300 bp (see example of results at the end of the manual). If many fragments larger than 500 bp are present , the best would be to contact your sequencing provider to ask what their requirements are, because it can vary depending on the sequencer. If you want to remove the large fragments you can use the size selection protocol described in the manual.<br /><br /></li>
<li><strong>What is the size of the adapters?</strong><br />The sum of the adapters is 128 bp.</li>
</ol>
</div>
<div class="extra-spaced">
<h2>MicroPlex Library Preparation Kit</h2>
<ol>
<li><strong>Can I use the available Illumina primers and validate them with the MicroPlex Kit v2?</strong><br /> Although the final flanking sequences of MicroPlex are the same as those used by Illumina, the PCR primers are not identical and part of them is supplied with the buffer. For this reason Illumina primers will not work as substitute.<br /><br /></li>
<li><strong>The BioAnalyzer profile of purified library shows the presence of low molecular weight peaks (primers/adaptors) in the samples. Should I re- purify the samples or they can be used directly to the sequencing? If the second purification is recommended, which ratio sample/AMPure beads should I use?</strong><br /> You can do a second round of purification using 1:1 ratio of AMPure beads to sample and this should get rid of the majority of the dimers.<br /><br /></li>
<li><strong>I am going to use the MicroPlex Library Preparation Kit v2 on ChIP samples . Our thermocycler has ramp rate 1.5°/s max while the protocol recommends using a ramp rate 3 to 5°/s. How would this affect the library prep?</strong><br /> We have not used a thermocycler with a ramp rate of 1.5 °C, which seems faster than most of thermocyclers. Too fast of a ramp rate may affect the primer annealing and ligation steps.<br /><br /></li>
<li><strong>What is the function of the replication stop site in the adapter loops?</strong><br /> The replication stop site in the adaptor loops function to stop the polymerase from continuing to copy the rest of the stem loop.<br /><br /></li>
<li><strong>I want to do ChIP-seq. Which ChIP-seq kit can I use for sample preparation prior to Microplex Library Preparation Kit v2?</strong><br /> In our portfolio there are several ChIP-seq kits compatible with Microplex Library Preparation Kit v2. Depending on your sample type and target studied you can use the following kits: iDeal ChIP-seq Kit for Transcription Factors (Cat. No. C01010055), iDeal ChIP-seq Kit for Histones (Cat. No. C01010051), True MicroChIP kit (Cat. No. C01010130), Universal Plant ChIP-seq Kit (Cat. No. C01010152). All these kits exist in manual and automated versions.<br /><br /></li>
<li><strong>Is Microplex Library Preparation Kit v2 compatible with exome enrichment methods?</strong><br /> Microplex Library Preparation Kit v2 is compatible with major exome and target enrichment products, including Agilent SureSelect<sup>®</sup>, Roche NimbleGen<sup>®</sup> SeqCap<sup>®</sup> EZ and custom panels.<br /><br /></li>
<li><strong>What is the nick that is mentioned in the kit method overview?</strong><br /> The nick is simply a gap between a stem adaptor and 3’ DNA end, as shown on the schema in the kit method overview.<br /><br /></li>
<li><strong>Are the indexes of the MicroPlex library preparation kit v2 located at i5 or i7?</strong><br /> The libraries generated with the MicroPlex kit v2 contain indices located at i7.<br /><br /></li>
<li><strong>Is there a need to use custom index read primers for the sequencing to read the 8nt iPCRtags?</strong><br /> There is no need for using custom Sequencing primer to sequence MicroPlex libraires. MicroPlex libraries can be sequenced using standard Illumina Sequencing kits and protocols.<br /><br /></li>
<li><strong>What is the advantage of using stem-loop adapter in the MicroPlex kit?</strong><br /> There are several advantages of using stem-loop adaptors. First of all, stem-loop adaptors prevent from self-ligation thus increases the ligation efficiency between the adapter and DNA fragment. Moreover, the background is reduced using ds adaptors with no single-stranded tails. Finally, adaptor-adaptor ligation is reduced using blocked 5’ ends.<br /><br /></li>
</ol>
</div>
<div class="extra-spaced">
<h2>IDeal Library Preparation Kit</h2>
<ol>
<li><strong>Are the index from the iDeal library Prep kit compatible with the MicroPlex library prep kit?</strong><br /> No, it is important to use only the indexes provided in the MicroPlex kit to ensure proper library preparation with this kit</li>
</ol>
</div>
</div>
</div>
</div>
</div>
</div>
</div>
<script>// <![CDATA[
$(document).ready(function() {
$('ul.tips-menu li a:first').trigger('focus');
});
// ]]></script>',
'no_promo' => false,
'in_menu' => true,
'online' => true,
'tabular' => true,
'hide' => true,
'all_format' => false,
'is_antibody' => false,
'slug' => 'library-preparation-for-ChIP-seq',
'cookies_tag_id' => null,
'meta_keywords' => 'Library preparation,Next Generation Sequencing,DNA fragments,MicroPlex Library,iDeal Library',
'meta_description' => 'Diagenode Kits have been Optimized for DNA library Preparation used for Next Generation Sequencing for a range of inputs.',
'meta_title' => 'DNA Library Preparation Kits,Microplex library Preparation Kits | Diagenode',
'modified' => '2021-12-20 14:30:28',
'created' => '2016-10-21 02:39:19',
'ProductsCategory' => array(
[maximum depth reached]
),
'CookiesTag' => array([maximum depth reached])
)
),
'Document' => array(
(int) 0 => array(
'id' => '1056',
'name' => 'Primer indexes for MicroPlex kit v3',
'description' => '<p>High Performance Library Preparation for Illumina<sup>®</sup> NGS Platforms</p>',
'image_id' => null,
'type' => 'Manual',
'url' => 'files/products/kits/Primer-indexes-for-Microplex-v3.pdf',
'slug' => 'primer-indexes-for-microplex-v3-manual',
'meta_keywords' => '',
'meta_description' => '',
'modified' => '2020-10-22 12:03:45',
'created' => '2019-07-18 11:53:57',
'ProductsDocument' => array(
[maximum depth reached]
)
)
),
'Feature' => array(),
'Image' => array(
(int) 0 => array(
'id' => '1759',
'name' => 'product/kits/chip-kit-icon.png',
'alt' => 'chip kit icon',
'modified' => '2018-01-18 15:26:13',
'created' => '2018-01-18 15:08:38',
'ProductsImage' => array(
[maximum depth reached]
)
)
),
'Promotion' => array(),
'Protocol' => array(),
'Publication' => array(),
'Testimonial' => array(),
'Area' => array(),
'SafetySheet' => array()
)
$country = 'US'
$countries_allowed = array(
(int) 0 => 'CA',
(int) 1 => 'US',
(int) 2 => 'IE',
(int) 3 => 'GB',
(int) 4 => 'DK',
(int) 5 => 'NO',
(int) 6 => 'SE',
(int) 7 => 'FI',
(int) 8 => 'NL',
(int) 9 => 'BE',
(int) 10 => 'LU',
(int) 11 => 'FR',
(int) 12 => 'DE',
(int) 13 => 'CH',
(int) 14 => 'AT',
(int) 15 => 'ES',
(int) 16 => 'IT',
(int) 17 => 'PT'
)
$outsource = false
$other_formats = array()
$edit = ''
$testimonials = ''
$featured_testimonials = ''
$related_products = '<li>
<div class="row">
<div class="small-12 columns">
<a href="/en/p/microplex-lib-prep-kit-v3-48-rxns"><img src="/img/product/kits/chip-kit-icon.png" alt="chip kit icon" class="th"/></a> </div>
<div class="small-12 columns">
<div class="small-6 columns" style="padding-left:0px;padding-right:0px;margin-top:-6px;margin-left:-1px">
<span class="success label" style="">C05010001</span>
</div>
<div class="small-6 columns text-right" style="padding-left:0px;padding-right:0px;margin-top:-6px">
<!--a href="#" style="color:#B21329"><i class="fa fa-info-circle"></i></a-->
<!-- BEGIN: ADD TO CART MODAL --><div id="cartModal-3032" class="reveal-modal small" data-reveal aria-labelledby="modalTitle" aria-hidden="true" role="dialog">
<form action="/en/carts/add/3032" id="CartAdd/3032Form" method="post" accept-charset="utf-8"><div style="display:none;"><input type="hidden" name="_method" value="POST"/></div><input type="hidden" name="data[Cart][product_id]" value="3032" id="CartProductId"/>
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p>Add <input name="data[Cart][quantity]" placeholder="1" value="1" min="1" style="width:60px;display:inline" type="number" id="CartQuantity" required="required"/> <strong> MicroPlex Library Preparation Kit v3 /48 rxns</strong> to my shopping cart.</p>
<div class="row">
<div class="small-6 medium-6 large-6 columns">
<button class="alert small button expand" onclick="$(this).addToCart('MicroPlex Library Preparation Kit v3 /48 rxns',
'C05010001',
'1555',
$('#CartQuantity').val());" name="checkout" id="checkout" value="checkout" type="submit">Checkout</button> </div>
<div class="small-6 medium-6 large-6 columns">
<button class="alert small button expand" onclick="$(this).addToCart('MicroPlex Library Preparation Kit v3 /48 rxns',
'C05010001',
'1555',
$('#CartQuantity').val());" name="keepshop" id="keepshop" type="submit">Keep shopping</button> </div>
</div>
</div>
</div>
</form><a class="close-reveal-modal" aria-label="Close">×</a></div><!-- END: ADD TO CART MODAL --><a href="#" id="microplex-lib-prep-kit-v3-48-rxns" data-reveal-id="cartModal-3032" class="" style="color:#B21329"><i class="fa fa-cart-plus"></i></a>
</div>
</div>
<div class="small-12 columns" >
<h6 style="height:60px">MicroPlex Library Preparation Kit v3 /48 rxns</h6>
</div>
</div>
</li>
'
$related = array(
'id' => '3032',
'antibody_id' => null,
'name' => 'MicroPlex Library Preparation Kit v3 /48 rxns',
'description' => '<p><a href="https://www.diagenode.com/files/products/kits/Microplex-library-prep-v3.pdf"><img src="https://www.diagenode.com/img/buttons/bt-manual.png" /></a></p>
<p>Diagenode’s <strong>MicroPlex Library Preparation Kits v3</strong> have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg. The kit MicroPlex v3 includes all buffers and enzymes necessary for the library preparation. For flexibility of the choice different formats of compatible primer indexes are available separately:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2">C05010005 - 96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p style="padding-left: 30px;">NEW! Unique dual indexes :</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1">C05010008 - 24 UDI for MicroPlex Kit v3 - Set I</a></li>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2">C05010009 - 24 UDI for MicroPlex Kit v3 - Set II</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong> with 24 dual indexes (8 nt)</li>
<li><strong>Validated with the IP-<a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">Star<sup>®</sup></a></strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit"> Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img alt="MicroPlex Library Preparation Kit v3 /48 rxns" src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label2' => '',
'info2' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label3' => '',
'info3' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'format' => '48 rxns',
'catalog_number' => 'C05010001',
'old_catalog_number' => '',
'sf_code' => 'C05010001-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1750',
'price_USD' => '1555',
'price_GBP' => '1555',
'price_JPY' => '286740',
'price_CNY' => '',
'price_AUD' => '3888',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => 'microplex-lib-prep-kit-v3-48-rxns',
'meta_title' => 'MicroPlex Library Preparation Kit v3 /48 rxns | Diagenode',
'meta_keywords' => 'MicroPlex Library Preparation Kit',
'meta_description' => 'MicroPlex Library Preparation Kits v3 have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg.',
'modified' => '2023-04-06 12:08:35',
'created' => '2019-07-16 16:03:58',
'ProductsRelated' => array(
'id' => '4541',
'product_id' => '3027',
'related_id' => '3032'
),
'Image' => array(
(int) 0 => array(
'id' => '1759',
'name' => 'product/kits/chip-kit-icon.png',
'alt' => 'chip kit icon',
'modified' => '2018-01-18 15:26:13',
'created' => '2018-01-18 15:08:38',
'ProductsImage' => array(
[maximum depth reached]
)
)
)
)
$rrbs_service = array(
(int) 0 => (int) 1894,
(int) 1 => (int) 1895
)
$chipseq_service = array(
(int) 0 => (int) 2683,
(int) 1 => (int) 1835,
(int) 2 => (int) 1836,
(int) 3 => (int) 2684,
(int) 4 => (int) 1838,
(int) 5 => (int) 1839,
(int) 6 => (int) 1856
)
$labelize = object(Closure) {
}
$old_catalog_number = ''
$country_code = 'US'
$label = '<img src="/img/banners/banner-customizer-back.png" alt=""/>'
$document = array(
'id' => '1056',
'name' => 'Primer indexes for MicroPlex kit v3',
'description' => '<p>High Performance Library Preparation for Illumina<sup>®</sup> NGS Platforms</p>',
'image_id' => null,
'type' => 'Manual',
'url' => 'files/products/kits/Primer-indexes-for-Microplex-v3.pdf',
'slug' => 'primer-indexes-for-microplex-v3-manual',
'meta_keywords' => '',
'meta_description' => '',
'modified' => '2020-10-22 12:03:45',
'created' => '2019-07-18 11:53:57',
'ProductsDocument' => array(
'id' => '2775',
'product_id' => '3027',
'document_id' => '1056'
)
)include - APP/View/Products/view.ctp, line 755
View::_evaluate() - CORE/Cake/View/View.php, line 971
View::_render() - CORE/Cake/View/View.php, line 933
View::render() - CORE/Cake/View/View.php, line 473
Controller::render() - CORE/Cake/Controller/Controller.php, line 963
ProductsController::slug() - APP/Controller/ProductsController.php, line 1052
ReflectionMethod::invokeArgs() - [internal], line ??
Controller::invokeAction() - CORE/Cake/Controller/Controller.php, line 491
Dispatcher::_invoke() - CORE/Cake/Routing/Dispatcher.php, line 193
Dispatcher::dispatch() - CORE/Cake/Routing/Dispatcher.php, line 167
[main] - APP/webroot/index.php, line 118
Notice (8): Undefined variable: message [APP/View/Products/view.ctp, line 755]Code Context<!-- BEGIN: REQUEST_FORM MODAL -->
<div id="request_formModal" class="reveal-modal medium" data-reveal aria-labelledby="modalTitle" aria-hidden="true" role="dialog">
<?= $this->element('Forms/simple_form', array('solution_of_interest' => $solution_of_interest, 'header' => $header, 'message' => $message, 'campaign_id' => $campaign_id)) ?>
$viewFile = '/home/website-server/www/app/View/Products/view.ctp'
$dataForView = array(
'language' => 'en',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'product' => array(
'Product' => array(
'id' => '3027',
'antibody_id' => null,
'name' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'description' => '<p>Diagenode’s <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns">MicroPlex Library Preparation Kits v3</a> have been extensively validated for ChIP-seq library prep from ChIP-derived DNA. The kit MicroPlex v3 has to be purchased with the compatible set of dual indexes. This set of dual indexes allows for multiplexing up to 96 samples.</p>
<p>Check all available sets of dual indexes:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong></li>
<li><strong>Validated with the </strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">IP-Star Compact Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>',
'label2' => '',
'info2' => '',
'label3' => '',
'info3' => '',
'format' => '96 rxns',
'catalog_number' => 'C05010005',
'old_catalog_number' => '',
'sf_code' => 'C05010005-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1355',
'price_USD' => '1300',
'price_GBP' => '1235',
'price_JPY' => '222015',
'price_CNY' => '',
'price_AUD' => '3250',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => '96-dual-indexes-for-microplex-kit-v3-set-2',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'modified' => '2020-10-23 17:11:41',
'created' => '2019-07-16 15:48:28',
'locale' => 'eng'
),
'Antibody' => array(
'host' => '*****',
'id' => null,
'name' => null,
'description' => null,
'clonality' => null,
'isotype' => null,
'lot' => null,
'concentration' => null,
'reactivity' => null,
'type' => null,
'purity' => null,
'classification' => null,
'application_table' => null,
'storage_conditions' => null,
'storage_buffer' => null,
'precautions' => null,
'uniprot_acc' => null,
'slug' => null,
'meta_keywords' => null,
'meta_description' => null,
'modified' => null,
'created' => null,
'select_label' => null
),
'Slave' => array(),
'Group' => array(),
'Related' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Application' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Category' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Document' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Feature' => array(),
'Image' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Promotion' => array(),
'Protocol' => array(),
'Publication' => array(),
'Testimonial' => array(),
'Area' => array(),
'SafetySheet' => array()
)
)
$language = 'en'
$meta_keywords = ''
$meta_description = '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns'
$meta_title = '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns'
$product = array(
'Product' => array(
'id' => '3027',
'antibody_id' => null,
'name' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'description' => '<p>Diagenode’s <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns">MicroPlex Library Preparation Kits v3</a> have been extensively validated for ChIP-seq library prep from ChIP-derived DNA. The kit MicroPlex v3 has to be purchased with the compatible set of dual indexes. This set of dual indexes allows for multiplexing up to 96 samples.</p>
<p>Check all available sets of dual indexes:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong></li>
<li><strong>Validated with the </strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">IP-Star Compact Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>',
'label2' => '',
'info2' => '',
'label3' => '',
'info3' => '',
'format' => '96 rxns',
'catalog_number' => 'C05010005',
'old_catalog_number' => '',
'sf_code' => 'C05010005-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1355',
'price_USD' => '1300',
'price_GBP' => '1235',
'price_JPY' => '222015',
'price_CNY' => '',
'price_AUD' => '3250',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => '96-dual-indexes-for-microplex-kit-v3-set-2',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'modified' => '2020-10-23 17:11:41',
'created' => '2019-07-16 15:48:28',
'locale' => 'eng'
),
'Antibody' => array(
'host' => '*****',
'id' => null,
'name' => null,
'description' => null,
'clonality' => null,
'isotype' => null,
'lot' => null,
'concentration' => null,
'reactivity' => null,
'type' => null,
'purity' => null,
'classification' => null,
'application_table' => null,
'storage_conditions' => null,
'storage_buffer' => null,
'precautions' => null,
'uniprot_acc' => null,
'slug' => null,
'meta_keywords' => null,
'meta_description' => null,
'modified' => null,
'created' => null,
'select_label' => null
),
'Slave' => array(),
'Group' => array(),
'Related' => array(
(int) 0 => array(
'id' => '3032',
'antibody_id' => null,
'name' => 'MicroPlex Library Preparation Kit v3 /48 rxns',
'description' => '<p><a href="https://www.diagenode.com/files/products/kits/Microplex-library-prep-v3.pdf"><img src="https://www.diagenode.com/img/buttons/bt-manual.png" /></a></p>
<p>Diagenode’s <strong>MicroPlex Library Preparation Kits v3</strong> have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg. The kit MicroPlex v3 includes all buffers and enzymes necessary for the library preparation. For flexibility of the choice different formats of compatible primer indexes are available separately:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2">C05010005 - 96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p style="padding-left: 30px;">NEW! Unique dual indexes :</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1">C05010008 - 24 UDI for MicroPlex Kit v3 - Set I</a></li>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2">C05010009 - 24 UDI for MicroPlex Kit v3 - Set II</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong> with 24 dual indexes (8 nt)</li>
<li><strong>Validated with the IP-<a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">Star<sup>®</sup></a></strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit"> Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img alt="MicroPlex Library Preparation Kit v3 /48 rxns" src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label2' => '',
'info2' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label3' => '',
'info3' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'format' => '48 rxns',
'catalog_number' => 'C05010001',
'old_catalog_number' => '',
'sf_code' => 'C05010001-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1750',
'price_USD' => '1555',
'price_GBP' => '1555',
'price_JPY' => '286740',
'price_CNY' => '',
'price_AUD' => '3888',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => 'microplex-lib-prep-kit-v3-48-rxns',
'meta_title' => 'MicroPlex Library Preparation Kit v3 /48 rxns | Diagenode',
'meta_keywords' => 'MicroPlex Library Preparation Kit',
'meta_description' => 'MicroPlex Library Preparation Kits v3 have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg.',
'modified' => '2023-04-06 12:08:35',
'created' => '2019-07-16 16:03:58',
'ProductsRelated' => array(
[maximum depth reached]
),
'Image' => array(
[maximum depth reached]
)
)
),
'Application' => array(
(int) 0 => array(
'id' => '14',
'position' => '10',
'parent_id' => '3',
'name' => 'DNA/RNA library preparation',
'description' => '<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p><span style="font-weight: 400;">Most of the major next-generation sequencing platforms require ligation of specific adaptor oligos to </span><a href="../applications/dna-rna-shearing"><span style="font-weight: 400;">fragmented DNA or RNA</span></a><span style="font-weight: 400;"> prior to sequencing</span></p>
<p><span style="font-weight: 400;">After input DNA has been fragmented, it is end-repaired and blunt-ended</span><span style="font-weight: 400;">. The next step is a A-tailing in which dAMP is added to the 3´ end of the blunt phosphorylated DNA fragments to prevent concatemerization and to allow the ligation of adaptors with complementary dT overhangs. In addition, barcoded adapters can be incorporated to facilitate multiplexing prior to or during amplification.</span></p>
<center><img src="https://www.diagenode.com/img/categories/library-prep/flux.png" /></center>
<p><span style="font-weight: 400;">Diagenode offers a comprehensive product portfolio for library preparation:<br /></span></p>
<strong><a href="https://www.diagenode.com/en/categories/Library-preparation-for-RNA-seq">D-Plex RNA-seq Library Preparation Kits</a></strong><br />
<p><span style="font-weight: 400;">Diagenode’s new RNA-sequencing solutions utilize the innovative c</span><span style="font-weight: 400;">apture and a</span><span style="font-weight: 400;">mplification by t</span><span style="font-weight: 400;">ailing and s</span><span style="font-weight: 400;">witching”</span><span style="font-weight: 400;">, a ligation-free method to produce DNA libraries for next generation sequencing from low input amounts of RNA. </span><span style="font-weight: 400;"></span><a href="../categories/Library-preparation-for-RNA-seq">Learn more</a></p>
<strong><a href="../categories/library-preparation-for-ChIP-seq">ChIP-seq and DNA sequencing library preparation solutions</a></strong><br />
<p><span style="font-weight: 400;">Our kits have been optimized for DNA library preparation used for next generation sequencing for a wide range of inputs. Using a simple three-step protocols, our</span><a href="http://www.diagenode.com/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns"><span style="font-weight: 400;"> </span></a><span style="font-weight: 400;">kits are an optimal choice for library preparation from DNA inputs down to 50 pg. </span><a href="../categories/library-preparation-for-ChIP-seq">Learn more</a></p>
<a href="../p/bioruptor-pico-sonication-device"><span style="font-weight: 400;"></span><strong>Bioruptor Pico - short fragments</strong></a><a href="../categories/library-preparation-for-ChIP-seq-and-DNA-sequencing"><span style="font-weight: 400;"></span></a><br />
<p><span style="font-weight: 400;"></span><span style="font-weight: 400;">Our well-cited Bioruptor Pico is the shearing device of choice for chromatin and DNA fragmentation. Obtain uniform and tight fragment distributions between 150bp -2kb. </span><a href="../p/bioruptor-pico-sonication-device">Learn more</a></p>
<strong><a href="../p/megaruptor2-1-unit"><span href="../p/bioruptor-pico-sonication-device">Megaruptor</span>® - long fragments</a></strong><a href="../p/bioruptor-pico-sonication-device"><span style="font-weight: 400;"></span></a><a href="../categories/library-preparation-for-ChIP-seq-and-DNA-sequencing"><span style="font-weight: 400;"></span></a><br />
<p><span style="font-weight: 400;"></span><span style="font-weight: 400;">The Megaruptor is designed to shear DNA from 3kb-75kb for long-read sequencing. <a href="../p/megaruptor2-1-unit">Learn more</a></span></p>
<span href="../p/bioruptor-pico-sonication-device"></span><span style="font-weight: 400;"></span></div>
</div>',
'in_footer' => false,
'in_menu' => true,
'online' => true,
'tabular' => true,
'slug' => 'library-preparation',
'meta_keywords' => 'Library preparation,Next Generation Sequencing,DNA fragments,MicroPlex Library,iDeal Library',
'meta_description' => 'Diagenode offers a comprehensive product portfolio for library preparation such as CATS RNA-seq Library Preparation Kits,ChIP-seq and DNA sequencing library preparation solutions',
'meta_title' => 'Library preparation for Next Generation Sequencing (NGS) | Diagenode',
'modified' => '2021-03-10 03:27:16',
'created' => '2014-08-16 07:47:56',
'ProductsApplication' => array(
[maximum depth reached]
)
)
),
'Category' => array(
(int) 0 => array(
'id' => '124',
'position' => '1',
'parent_id' => '15',
'name' => 'Library preparation for ChIP-seq',
'description' => '<div class="row">
<div class="large-12 columns text-justify">
<p>Library preparation following ChIP can be challenging due to the limited amount of DNA recovered after ChIP. Diagenode has developed the optimal solutions for ChIP-seq using two different approaches: the ligation-based library preparation on purified DNA or the tagmentation-based ChIPmentation.</p>
</div>
</div>
<div class="row extra-spaced">
<div class="large-12 columns"><center><a href="https://www.diagenode.com/en/pages/form-microplex-promo" target="_blank"></a></center></div>
</div>
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div id="portal" class="main-portal">
<div class="portal-inner"><nav class="portal-nav">
<ul data-tab="" class="tips-menu">
<li><a href="#panel1" class="tips portal button">Ligation-based library prep</a></li>
<li><a href="#panel2" class="tips portal button">ChIPmentation</a></li>
<li><a href="#panel3" class="tips portal button">Kit choice guide</a></li>
<li><a href="#panel4" class="tips portal button">Resources</a></li>
<li><a href="#panel5" class="tips portal button">FAQs</a></li>
</ul>
</nav></div>
</div>
<div class="tabs-content">
<div class="content active" id="panel1">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v5" style="color: #13b29c;"><i class="fa fa-caret-right"></i> Standard input library prep</a>
<div id="v5" class="content">
<div class="small-12 medium-12 large-12 columns">
<p>The <strong>iDeal Library Preparation Kit</strong> reliably converts DNA into indexed libraries for next-generation sequencing, with input amounts down to <strong>5 ng</strong>. Our kit offers a simple and fast workflow, high yields, and ready-to-sequence DNA on the Illumina platform.</p>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> <strong>Sample</strong>: Fragmented dsDNA</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Input</strong>: 5 ng – 1 µg</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Fast protocol</strong>: 3 hours</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Easy processing</strong>: 3 steps</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Indexing</strong>: single indexes for multiplexing up to 24 samples</li>
<li><i class="fa fa-arrow-circle-right"></i> Manual and automated protocols available</li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<ul class="square">
<li>MeDIP-seq library prep</li>
<li>Genomic DNA sequencing</li>
<li>High input ChIP-seq</li>
</ul>
</div>
<div class="extra-spaced">
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010020</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" style="color: #b21329;" target="_blank">iDeal Library Preparation Kit x24 (incl. Index Primer Set 1)</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010021</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" style="color: #b21329;" target="_blank">Index Primer Set 2 (iDeal Lib. Prep Kit x24)</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
</li>
</ul>
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v4" style="color: #13b29c;"><i class="fa fa-caret-right"></i> Low input library prep</a>
<div id="v4" class="content active"><center><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" target="_blank"><img src="https://www.diagenode.com/img/banners/banner-microplex-v3-580.jpg" class="extra-spaced" /></a></center>
<div align="center"><a href="https://www.diagenode.com/pages/form-microplex3" class="center alert radius button extra-spaced"><i class="fa fa-info"></i> Contact us</a></div>
<div class="extra-spaced">
<p>Diagenode’s <strong>MicroPlex Library Preparation kits</strong> have been extensively validated for ChIP-seq samples. Generated libraries are compatible with single-end or paired-end sequencing. MicroPlex chemistry (using stem-loop adapters ) is specifically developed and optimized to generate DNA libraries with high molecular complexity from the lowest input amounts. Only <strong>50 pg to 50 ng</strong> of fragmented double-stranded DNA is required for library preparation. The entire <strong>three-step workflow</strong> takes place in a <strong>single tube</strong> or well in about <strong>2 hours</strong>. No intermediate purification steps and no sample transfers are necessary to prevent handling errors and loss of valuable samples.</p>
</div>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> <strong>Sample</strong>: Fragmented dsDNA</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Low input</strong>: 50 pg – 50 ng</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Fast protocol</strong>: 2 hours</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Easy processing</strong>: 3 steps in 1 tube</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>No intermediate purification</strong></li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
<li><i class="fa fa-arrow-circle-right"></i> Manual and automated protocols available</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<ul class="square">
<li>ChIP-seq library prep from ChIP-derived DNA</li>
<li>Low input DNA sequencing</li>
</ul>
</div>
<h2>Two versions are available:</h2>
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v2" style="color: #13b29c;"><i class="fa fa-caret-right"></i> MicroPlex Library Preparation Kit v2 with single indexes</a>
<div id="v2" class="content">
<p>The MicroPlex Library Preparation Kit v2 contains all necessary reagents including single indexes for multiplexing up to 48 samples using single barcoding.</p>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010012</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v2 (12 indexes)</a></td>
<td class="format">12 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</li>
<li class="accordion-navigation"><a href="#v3" style="color: #13b29c;"><i class="fa fa-caret-right"></i> MicroPlex Library Preparation Kit v3 with dual indexes <strong><span class="diacol">NEW!</span></strong></a>
<div id="v3" class="content active">
<p>In this version the library preparation reagents and the dual indexes are available separately allowing for the flexibility choosing the number of indexes. MicroPlex v3 has multiplexing capacities up to 384 samples.</p>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010001</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v3 /48 rxns</a></td>
<td class="format">48 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010002</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v3 /96 rxns</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
<h4>DUAL INDEXES</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010003</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" style="color: #b21329;" target="_blank">24 Dual indexes for MicroPlex Kit v3</a></td>
<td class="format">48 rxns</td>
<td><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010004</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set I</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010005</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set II</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010006</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set III</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010007</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set IV</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</li>
</ul>
</div>
</li>
</ul>
</div>
</div>
</div>
<div class="content active" id="panel2">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<p>The TAG Kit for ChIPmentation offers an optimized ChIP-seq library preparation solution based on tagmentation. This kit includes reagents for tagmentation-based library preparation integrated in the IP and is compatible with any ChIP protocol based on magnetic beads. The primer indexes for multiplexing must be purchased separately and are available as a reference: <a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" target="_blank">24 SI for ChIPmentation</a>, Cat. No. C01011031. Alternatively, for histone marks, Diagenode proposes the complete solution (including all buffers for ChIP, tagmentation and multiplexing): <a href="https://www.diagenode.com/en/p/manual-chipmentation-kit-for-histones-24-rxns" target="_blank">ChIPmentation for Histones</a>.</p>
</div>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> Sample: chromatin-antibody-magnetic beads complexes</li>
<li><i class="fa fa-arrow-circle-right"></i> Input: chromatin from 5 K – 4 M cells</li>
<li><i class="fa fa-arrow-circle-right"></i> Easy and fast protocol</li>
<li><i class="fa fa-arrow-circle-right"></i> Compatible with any ChIP protocol based on magnetic beads</li>
<li><i class="fa fa-arrow-circle-right"></i> No adapter dimers</li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<p class="lead"><em><strong>TAG kit for ChIPmentation</strong></em></p>
<ul class="square">
<li>ChIPmentation library preparation</li>
</ul>
<p class="lead"><em><strong>24 SI for for ChIPmentation</strong></em></p>
<ul class="square">
<li>ChIPmentation library preparation</li>
<li>Tagmentation-based library preparation methods like ATAC-seq, CUT&Tag</li>
</ul>
</div>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C01011030</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" style="color: #b21329;" target="_blank">TAG Kit for ChIPmentation</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C01011031</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" style="color: #b21329;" target="_blank">24 SI for ChIPmentation</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
<div class="content" id="panel3">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<h3 class="text-center diacol"><em>How to choose your library preparation kit?</em></h3>
</div>
<table class="noborder">
<tbody>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Sample</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Chromatin-antibody-beads complex</p>
</td>
<td colspan="2">
<p class="text-center" style="font-size: 15px;">Purified DNA</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Purified DNA</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td colspan="2"><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Application</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">ChIPmentation</p>
</td>
<td colspan="2">
<p class="text-center" style="font-size: 15px;">ChIP-seq library prep<br /> Low input DNA sequencing</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">MeDIP-seq library prep<br /> Genomic DNA sequencing<br /> High input ChIP-seq</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td colspan="2"><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Input</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Chromatin: 5 K to 4 M cells</p>
</td>
<td colspan="2"">
<p class="text-center" style="font-size: 15px;">DNA: 50 pg – 50 ng</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">DNA: 5 ng – 1 µg</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow-45-left.png" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow-45-right.png" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Multiplexing</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 24 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 384 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 48 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 24 samples</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Indexes</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Dual indexes (DI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Kit</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>TAG Kit for ChIPmentation</strong><br /> (indexes not included in the kit)</p>
<p class="text-center"><strong>Kit</strong><br /> <a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" target="_blank">C01011030 – 24 rxns</a></p>
<p class="text-center"><strong>Single indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" target="_blank">C01011031 – 24 SI/24 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>MicroPlex Library Preparation Kit v3</strong><br />(dual indexes not included in the kit)</p>
<p class="text-center"><strong>Kit</strong><br /> <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" target="_blank">C05010001 - 48 rxns</a><br /> <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" target="_blank">C05010002 - 96 rxns</a></p>
<br />
<p class="text-center"><strong>Unique dual indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1" target="_blank">C05010008 - Set I 24 UDI / 24 rxns</a><br /> <a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2" target="_blank">C05010009 - Set II 24 UDI/ 24 rxns</a></p>
<p class="text-center"><strong>Dual indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" target="_blank">C05010003 - 24 DI/ 48 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" target="_blank">C05010004 - Set I 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" target="_blank">C05010005 - Set II 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" target="_blank">C05010006 - Set III 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" target="_blank">C05010007 - Set IV 96 DI/ 96 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>MicroPlex Library Preparation Kit v2</strong><br />(single indexes included in the kit)</p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" target="_blank">C05010012 - 12 SI/ 12 rxns</a><br /> <a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x48-12-indices-48-rxns" target="_blank">C05010013 - 12 SI/ 48 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>iDeal Library Preparation Kit</strong><br />(Set 1 of indexes included in the kit)</p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" target="_blank">C05010020 - 12 SI/ 24 rxns</a></p>
<p class="text-center" style="font-size: 15px;"><strong>Index Primer Set 2</strong></p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" target="_blank">C05010021 - 12 SI/ 24 rxns</a></p>
</td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
<div class="content" id="panel4">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p>Combined chromatin immunoprecipitation and next-generation sequencing (ChIP-seq) has become the gold standard to investigate genome-wide epigenetic profiles. However, ChIP from a limited amount of cells has been a challenge. Here we provide a complete and robust workflow solution for successful ChIP-seq from small numbers of cells using the True MicroChIP kit and MicroPlex Library Preparation kit.</p>
<blockquote><span class="label-green" style="margin-bottom: 16px; margin-left: -22px;">APPLICATION NOTE</span>
<div class="row">
<div class="small-12 medium-6 large-6 columns"><center><img src="https://www.diagenode.com/img/categories/microplex/chip-efficiency-on-10000-cells.jpg" /></center>
<p><small><em>ChIP efficiency on 10,000 cells</em></small></p>
</div>
<div class="small-12 medium-6 large-6 columns">
<p><strong>From minuscule amounts to magnificent results:</strong><br /> reliable ChIP-seq data from 10,000 cells with the True MicroChIP™ and the MicroPlex Library Preparation™ kits.</p>
<a href="https://www.diagenode.com/files/application_notes/True_MicroChIP_and_MicroPlex_kits_Application_Note.pdf" class="details small button" target="_blank">DOWNLOAD</a></div>
</div>
</blockquote>
<blockquote><span class="label-green" style="margin-bottom: 16px; margin-left: -22px;">APPLICATION NOTE</span>
<div class="row">
<div class="small-12 medium-6 large-6 columns"><center><img src="https://www.diagenode.com/img/categories/microplex/quality-control-check.jpg" /></center>
<p class="text-left"><small><em>Quality control check of a ChIP-seq library on the Fragment Analyzer. High Efficiency ChIP performed on 10,000 cells</em></small></p>
</div>
<div class="small-12 medium-6 large-6 columns">
<p class="text-left"><strong>Best Workflow Practices for ChIP-seq Analysis with Small Samples</strong></p>
<a href="https://www.diagenode.com/files/application_notes/Diagenode_AATI_Joint.pdf" class="details small button" target="_blank">DOWNLOAD</a></div>
</div>
</blockquote>
</div>
</div>
</div>
<div class="content" id="panel5">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<h2>TAG Kit for ChIPmentation</h2>
<ol>
<li><strong>What is the difference between tagmentation and ChIPmentation?</strong><br />Tagmentation is a reaction where an enzyme (a transposase) cleaves DNA and incorporates sequencing adaptors at the ends of the fragments in one step. In our ChIPmentation technology we combine chromatin immunoprecipitation and tagmentation in one streamlined workflow where the tagmentation step occurs directly on chromatin.<br /><br /></li>
<li><strong>What is the expected concentration of ChIPmentation libraries?</strong><br />The concentration of libraries that you need to reach will depend on the sensitivity of the machine and kits that you will use to perform the quality control and the sequencing of your libraries. Usually a concentration of 4-8 ng/μl is enough for a quality control using the Qubit High Sensitivity assay (ThermoFischer Scientific) and the High Sensitivity chip for BioAnalyzer (Agilent) and for sequencing on Illumina HiSeq3000/4000.<br /><br /></li>
<li><strong>Does the ChIPmentation approach work on plants?</strong><br />Our ChIPmentation solution has been validated on human cells and we do not have any data on plants. It should be compatible. We would recommend using our Universal Plant ChIP Kit in combination with the TAG Kit for ChIPmentation and the 24 SI for ChIPmentation.<br /><br /></li>
<li><strong>What is the size of the fragments after the tagmentation?</strong><br />The size of the fragments at the end of the ChIPmentation protocol can vary depending on many parameters like the shearing efficiency, the antibody used or the tagmentation time. However, with our standard protocol we usually obtain a library peak which is around 200-300 bp (see example of results at the end of the manual). If many fragments larger than 500 bp are present , the best would be to contact your sequencing provider to ask what their requirements are, because it can vary depending on the sequencer. If you want to remove the large fragments you can use the size selection protocol described in the manual.<br /><br /></li>
<li><strong>What is the size of the adapters?</strong><br />The sum of the adapters is 128 bp.</li>
</ol>
</div>
<div class="extra-spaced">
<h2>MicroPlex Library Preparation Kit</h2>
<ol>
<li><strong>Can I use the available Illumina primers and validate them with the MicroPlex Kit v2?</strong><br /> Although the final flanking sequences of MicroPlex are the same as those used by Illumina, the PCR primers are not identical and part of them is supplied with the buffer. For this reason Illumina primers will not work as substitute.<br /><br /></li>
<li><strong>The BioAnalyzer profile of purified library shows the presence of low molecular weight peaks (primers/adaptors) in the samples. Should I re- purify the samples or they can be used directly to the sequencing? If the second purification is recommended, which ratio sample/AMPure beads should I use?</strong><br /> You can do a second round of purification using 1:1 ratio of AMPure beads to sample and this should get rid of the majority of the dimers.<br /><br /></li>
<li><strong>I am going to use the MicroPlex Library Preparation Kit v2 on ChIP samples . Our thermocycler has ramp rate 1.5°/s max while the protocol recommends using a ramp rate 3 to 5°/s. How would this affect the library prep?</strong><br /> We have not used a thermocycler with a ramp rate of 1.5 °C, which seems faster than most of thermocyclers. Too fast of a ramp rate may affect the primer annealing and ligation steps.<br /><br /></li>
<li><strong>What is the function of the replication stop site in the adapter loops?</strong><br /> The replication stop site in the adaptor loops function to stop the polymerase from continuing to copy the rest of the stem loop.<br /><br /></li>
<li><strong>I want to do ChIP-seq. Which ChIP-seq kit can I use for sample preparation prior to Microplex Library Preparation Kit v2?</strong><br /> In our portfolio there are several ChIP-seq kits compatible with Microplex Library Preparation Kit v2. Depending on your sample type and target studied you can use the following kits: iDeal ChIP-seq Kit for Transcription Factors (Cat. No. C01010055), iDeal ChIP-seq Kit for Histones (Cat. No. C01010051), True MicroChIP kit (Cat. No. C01010130), Universal Plant ChIP-seq Kit (Cat. No. C01010152). All these kits exist in manual and automated versions.<br /><br /></li>
<li><strong>Is Microplex Library Preparation Kit v2 compatible with exome enrichment methods?</strong><br /> Microplex Library Preparation Kit v2 is compatible with major exome and target enrichment products, including Agilent SureSelect<sup>®</sup>, Roche NimbleGen<sup>®</sup> SeqCap<sup>®</sup> EZ and custom panels.<br /><br /></li>
<li><strong>What is the nick that is mentioned in the kit method overview?</strong><br /> The nick is simply a gap between a stem adaptor and 3’ DNA end, as shown on the schema in the kit method overview.<br /><br /></li>
<li><strong>Are the indexes of the MicroPlex library preparation kit v2 located at i5 or i7?</strong><br /> The libraries generated with the MicroPlex kit v2 contain indices located at i7.<br /><br /></li>
<li><strong>Is there a need to use custom index read primers for the sequencing to read the 8nt iPCRtags?</strong><br /> There is no need for using custom Sequencing primer to sequence MicroPlex libraires. MicroPlex libraries can be sequenced using standard Illumina Sequencing kits and protocols.<br /><br /></li>
<li><strong>What is the advantage of using stem-loop adapter in the MicroPlex kit?</strong><br /> There are several advantages of using stem-loop adaptors. First of all, stem-loop adaptors prevent from self-ligation thus increases the ligation efficiency between the adapter and DNA fragment. Moreover, the background is reduced using ds adaptors with no single-stranded tails. Finally, adaptor-adaptor ligation is reduced using blocked 5’ ends.<br /><br /></li>
</ol>
</div>
<div class="extra-spaced">
<h2>IDeal Library Preparation Kit</h2>
<ol>
<li><strong>Are the index from the iDeal library Prep kit compatible with the MicroPlex library prep kit?</strong><br /> No, it is important to use only the indexes provided in the MicroPlex kit to ensure proper library preparation with this kit</li>
</ol>
</div>
</div>
</div>
</div>
</div>
</div>
</div>
<script>// <![CDATA[
$(document).ready(function() {
$('ul.tips-menu li a:first').trigger('focus');
});
// ]]></script>',
'no_promo' => false,
'in_menu' => true,
'online' => true,
'tabular' => true,
'hide' => true,
'all_format' => false,
'is_antibody' => false,
'slug' => 'library-preparation-for-ChIP-seq',
'cookies_tag_id' => null,
'meta_keywords' => 'Library preparation,Next Generation Sequencing,DNA fragments,MicroPlex Library,iDeal Library',
'meta_description' => 'Diagenode Kits have been Optimized for DNA library Preparation used for Next Generation Sequencing for a range of inputs.',
'meta_title' => 'DNA Library Preparation Kits,Microplex library Preparation Kits | Diagenode',
'modified' => '2021-12-20 14:30:28',
'created' => '2016-10-21 02:39:19',
'ProductsCategory' => array(
[maximum depth reached]
),
'CookiesTag' => array([maximum depth reached])
)
),
'Document' => array(
(int) 0 => array(
'id' => '1056',
'name' => 'Primer indexes for MicroPlex kit v3',
'description' => '<p>High Performance Library Preparation for Illumina<sup>®</sup> NGS Platforms</p>',
'image_id' => null,
'type' => 'Manual',
'url' => 'files/products/kits/Primer-indexes-for-Microplex-v3.pdf',
'slug' => 'primer-indexes-for-microplex-v3-manual',
'meta_keywords' => '',
'meta_description' => '',
'modified' => '2020-10-22 12:03:45',
'created' => '2019-07-18 11:53:57',
'ProductsDocument' => array(
[maximum depth reached]
)
)
),
'Feature' => array(),
'Image' => array(
(int) 0 => array(
'id' => '1759',
'name' => 'product/kits/chip-kit-icon.png',
'alt' => 'chip kit icon',
'modified' => '2018-01-18 15:26:13',
'created' => '2018-01-18 15:08:38',
'ProductsImage' => array(
[maximum depth reached]
)
)
),
'Promotion' => array(),
'Protocol' => array(),
'Publication' => array(),
'Testimonial' => array(),
'Area' => array(),
'SafetySheet' => array()
)
$country = 'US'
$countries_allowed = array(
(int) 0 => 'CA',
(int) 1 => 'US',
(int) 2 => 'IE',
(int) 3 => 'GB',
(int) 4 => 'DK',
(int) 5 => 'NO',
(int) 6 => 'SE',
(int) 7 => 'FI',
(int) 8 => 'NL',
(int) 9 => 'BE',
(int) 10 => 'LU',
(int) 11 => 'FR',
(int) 12 => 'DE',
(int) 13 => 'CH',
(int) 14 => 'AT',
(int) 15 => 'ES',
(int) 16 => 'IT',
(int) 17 => 'PT'
)
$outsource = false
$other_formats = array()
$edit = ''
$testimonials = ''
$featured_testimonials = ''
$related_products = '<li>
<div class="row">
<div class="small-12 columns">
<a href="/en/p/microplex-lib-prep-kit-v3-48-rxns"><img src="/img/product/kits/chip-kit-icon.png" alt="chip kit icon" class="th"/></a> </div>
<div class="small-12 columns">
<div class="small-6 columns" style="padding-left:0px;padding-right:0px;margin-top:-6px;margin-left:-1px">
<span class="success label" style="">C05010001</span>
</div>
<div class="small-6 columns text-right" style="padding-left:0px;padding-right:0px;margin-top:-6px">
<!--a href="#" style="color:#B21329"><i class="fa fa-info-circle"></i></a-->
<!-- BEGIN: ADD TO CART MODAL --><div id="cartModal-3032" class="reveal-modal small" data-reveal aria-labelledby="modalTitle" aria-hidden="true" role="dialog">
<form action="/en/carts/add/3032" id="CartAdd/3032Form" method="post" accept-charset="utf-8"><div style="display:none;"><input type="hidden" name="_method" value="POST"/></div><input type="hidden" name="data[Cart][product_id]" value="3032" id="CartProductId"/>
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p>Add <input name="data[Cart][quantity]" placeholder="1" value="1" min="1" style="width:60px;display:inline" type="number" id="CartQuantity" required="required"/> <strong> MicroPlex Library Preparation Kit v3 /48 rxns</strong> to my shopping cart.</p>
<div class="row">
<div class="small-6 medium-6 large-6 columns">
<button class="alert small button expand" onclick="$(this).addToCart('MicroPlex Library Preparation Kit v3 /48 rxns',
'C05010001',
'1555',
$('#CartQuantity').val());" name="checkout" id="checkout" value="checkout" type="submit">Checkout</button> </div>
<div class="small-6 medium-6 large-6 columns">
<button class="alert small button expand" onclick="$(this).addToCart('MicroPlex Library Preparation Kit v3 /48 rxns',
'C05010001',
'1555',
$('#CartQuantity').val());" name="keepshop" id="keepshop" type="submit">Keep shopping</button> </div>
</div>
</div>
</div>
</form><a class="close-reveal-modal" aria-label="Close">×</a></div><!-- END: ADD TO CART MODAL --><a href="#" id="microplex-lib-prep-kit-v3-48-rxns" data-reveal-id="cartModal-3032" class="" style="color:#B21329"><i class="fa fa-cart-plus"></i></a>
</div>
</div>
<div class="small-12 columns" >
<h6 style="height:60px">MicroPlex Library Preparation Kit v3 /48 rxns</h6>
</div>
</div>
</li>
'
$related = array(
'id' => '3032',
'antibody_id' => null,
'name' => 'MicroPlex Library Preparation Kit v3 /48 rxns',
'description' => '<p><a href="https://www.diagenode.com/files/products/kits/Microplex-library-prep-v3.pdf"><img src="https://www.diagenode.com/img/buttons/bt-manual.png" /></a></p>
<p>Diagenode’s <strong>MicroPlex Library Preparation Kits v3</strong> have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg. The kit MicroPlex v3 includes all buffers and enzymes necessary for the library preparation. For flexibility of the choice different formats of compatible primer indexes are available separately:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2">C05010005 - 96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p style="padding-left: 30px;">NEW! Unique dual indexes :</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1">C05010008 - 24 UDI for MicroPlex Kit v3 - Set I</a></li>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2">C05010009 - 24 UDI for MicroPlex Kit v3 - Set II</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong> with 24 dual indexes (8 nt)</li>
<li><strong>Validated with the IP-<a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">Star<sup>®</sup></a></strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit"> Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img alt="MicroPlex Library Preparation Kit v3 /48 rxns" src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label2' => '',
'info2' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label3' => '',
'info3' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'format' => '48 rxns',
'catalog_number' => 'C05010001',
'old_catalog_number' => '',
'sf_code' => 'C05010001-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1750',
'price_USD' => '1555',
'price_GBP' => '1555',
'price_JPY' => '286740',
'price_CNY' => '',
'price_AUD' => '3888',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => 'microplex-lib-prep-kit-v3-48-rxns',
'meta_title' => 'MicroPlex Library Preparation Kit v3 /48 rxns | Diagenode',
'meta_keywords' => 'MicroPlex Library Preparation Kit',
'meta_description' => 'MicroPlex Library Preparation Kits v3 have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg.',
'modified' => '2023-04-06 12:08:35',
'created' => '2019-07-16 16:03:58',
'ProductsRelated' => array(
'id' => '4541',
'product_id' => '3027',
'related_id' => '3032'
),
'Image' => array(
(int) 0 => array(
'id' => '1759',
'name' => 'product/kits/chip-kit-icon.png',
'alt' => 'chip kit icon',
'modified' => '2018-01-18 15:26:13',
'created' => '2018-01-18 15:08:38',
'ProductsImage' => array(
[maximum depth reached]
)
)
)
)
$rrbs_service = array(
(int) 0 => (int) 1894,
(int) 1 => (int) 1895
)
$chipseq_service = array(
(int) 0 => (int) 2683,
(int) 1 => (int) 1835,
(int) 2 => (int) 1836,
(int) 3 => (int) 2684,
(int) 4 => (int) 1838,
(int) 5 => (int) 1839,
(int) 6 => (int) 1856
)
$labelize = object(Closure) {
}
$old_catalog_number = ''
$country_code = 'US'
$label = '<img src="/img/banners/banner-customizer-back.png" alt=""/>'
$document = array(
'id' => '1056',
'name' => 'Primer indexes for MicroPlex kit v3',
'description' => '<p>High Performance Library Preparation for Illumina<sup>®</sup> NGS Platforms</p>',
'image_id' => null,
'type' => 'Manual',
'url' => 'files/products/kits/Primer-indexes-for-Microplex-v3.pdf',
'slug' => 'primer-indexes-for-microplex-v3-manual',
'meta_keywords' => '',
'meta_description' => '',
'modified' => '2020-10-22 12:03:45',
'created' => '2019-07-18 11:53:57',
'ProductsDocument' => array(
'id' => '2775',
'product_id' => '3027',
'document_id' => '1056'
)
)include - APP/View/Products/view.ctp, line 755
View::_evaluate() - CORE/Cake/View/View.php, line 971
View::_render() - CORE/Cake/View/View.php, line 933
View::render() - CORE/Cake/View/View.php, line 473
Controller::render() - CORE/Cake/Controller/Controller.php, line 963
ProductsController::slug() - APP/Controller/ProductsController.php, line 1052
ReflectionMethod::invokeArgs() - [internal], line ??
Controller::invokeAction() - CORE/Cake/Controller/Controller.php, line 491
Dispatcher::_invoke() - CORE/Cake/Routing/Dispatcher.php, line 193
Dispatcher::dispatch() - CORE/Cake/Routing/Dispatcher.php, line 167
[main] - APP/webroot/index.php, line 118
Notice (8): Undefined variable: campaign_id [APP/View/Products/view.ctp, line 755]Code Context<!-- BEGIN: REQUEST_FORM MODAL -->
<div id="request_formModal" class="reveal-modal medium" data-reveal aria-labelledby="modalTitle" aria-hidden="true" role="dialog">
<?= $this->element('Forms/simple_form', array('solution_of_interest' => $solution_of_interest, 'header' => $header, 'message' => $message, 'campaign_id' => $campaign_id)) ?>
$viewFile = '/home/website-server/www/app/View/Products/view.ctp'
$dataForView = array(
'language' => 'en',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'product' => array(
'Product' => array(
'id' => '3027',
'antibody_id' => null,
'name' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'description' => '<p>Diagenode’s <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns">MicroPlex Library Preparation Kits v3</a> have been extensively validated for ChIP-seq library prep from ChIP-derived DNA. The kit MicroPlex v3 has to be purchased with the compatible set of dual indexes. This set of dual indexes allows for multiplexing up to 96 samples.</p>
<p>Check all available sets of dual indexes:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong></li>
<li><strong>Validated with the </strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">IP-Star Compact Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>',
'label2' => '',
'info2' => '',
'label3' => '',
'info3' => '',
'format' => '96 rxns',
'catalog_number' => 'C05010005',
'old_catalog_number' => '',
'sf_code' => 'C05010005-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1355',
'price_USD' => '1300',
'price_GBP' => '1235',
'price_JPY' => '222015',
'price_CNY' => '',
'price_AUD' => '3250',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => '96-dual-indexes-for-microplex-kit-v3-set-2',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'modified' => '2020-10-23 17:11:41',
'created' => '2019-07-16 15:48:28',
'locale' => 'eng'
),
'Antibody' => array(
'host' => '*****',
'id' => null,
'name' => null,
'description' => null,
'clonality' => null,
'isotype' => null,
'lot' => null,
'concentration' => null,
'reactivity' => null,
'type' => null,
'purity' => null,
'classification' => null,
'application_table' => null,
'storage_conditions' => null,
'storage_buffer' => null,
'precautions' => null,
'uniprot_acc' => null,
'slug' => null,
'meta_keywords' => null,
'meta_description' => null,
'modified' => null,
'created' => null,
'select_label' => null
),
'Slave' => array(),
'Group' => array(),
'Related' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Application' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Category' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Document' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Feature' => array(),
'Image' => array(
(int) 0 => array(
[maximum depth reached]
)
),
'Promotion' => array(),
'Protocol' => array(),
'Publication' => array(),
'Testimonial' => array(),
'Area' => array(),
'SafetySheet' => array()
)
)
$language = 'en'
$meta_keywords = ''
$meta_description = '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns'
$meta_title = '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns'
$product = array(
'Product' => array(
'id' => '3027',
'antibody_id' => null,
'name' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'description' => '<p>Diagenode’s <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns">MicroPlex Library Preparation Kits v3</a> have been extensively validated for ChIP-seq library prep from ChIP-derived DNA. The kit MicroPlex v3 has to be purchased with the compatible set of dual indexes. This set of dual indexes allows for multiplexing up to 96 samples.</p>
<p>Check all available sets of dual indexes:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong></li>
<li><strong>Validated with the </strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">IP-Star Compact Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>',
'label2' => '',
'info2' => '',
'label3' => '',
'info3' => '',
'format' => '96 rxns',
'catalog_number' => 'C05010005',
'old_catalog_number' => '',
'sf_code' => 'C05010005-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1355',
'price_USD' => '1300',
'price_GBP' => '1235',
'price_JPY' => '222015',
'price_CNY' => '',
'price_AUD' => '3250',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => '96-dual-indexes-for-microplex-kit-v3-set-2',
'meta_title' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'meta_keywords' => '',
'meta_description' => '96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns',
'modified' => '2020-10-23 17:11:41',
'created' => '2019-07-16 15:48:28',
'locale' => 'eng'
),
'Antibody' => array(
'host' => '*****',
'id' => null,
'name' => null,
'description' => null,
'clonality' => null,
'isotype' => null,
'lot' => null,
'concentration' => null,
'reactivity' => null,
'type' => null,
'purity' => null,
'classification' => null,
'application_table' => null,
'storage_conditions' => null,
'storage_buffer' => null,
'precautions' => null,
'uniprot_acc' => null,
'slug' => null,
'meta_keywords' => null,
'meta_description' => null,
'modified' => null,
'created' => null,
'select_label' => null
),
'Slave' => array(),
'Group' => array(),
'Related' => array(
(int) 0 => array(
'id' => '3032',
'antibody_id' => null,
'name' => 'MicroPlex Library Preparation Kit v3 /48 rxns',
'description' => '<p><a href="https://www.diagenode.com/files/products/kits/Microplex-library-prep-v3.pdf"><img src="https://www.diagenode.com/img/buttons/bt-manual.png" /></a></p>
<p>Diagenode’s <strong>MicroPlex Library Preparation Kits v3</strong> have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg. The kit MicroPlex v3 includes all buffers and enzymes necessary for the library preparation. For flexibility of the choice different formats of compatible primer indexes are available separately:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2">C05010005 - 96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p style="padding-left: 30px;">NEW! Unique dual indexes :</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1">C05010008 - 24 UDI for MicroPlex Kit v3 - Set I</a></li>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2">C05010009 - 24 UDI for MicroPlex Kit v3 - Set II</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong> with 24 dual indexes (8 nt)</li>
<li><strong>Validated with the IP-<a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">Star<sup>®</sup></a></strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit"> Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img alt="MicroPlex Library Preparation Kit v3 /48 rxns" src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label2' => '',
'info2' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label3' => '',
'info3' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'format' => '48 rxns',
'catalog_number' => 'C05010001',
'old_catalog_number' => '',
'sf_code' => 'C05010001-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1750',
'price_USD' => '1555',
'price_GBP' => '1555',
'price_JPY' => '286740',
'price_CNY' => '',
'price_AUD' => '3888',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => 'microplex-lib-prep-kit-v3-48-rxns',
'meta_title' => 'MicroPlex Library Preparation Kit v3 /48 rxns | Diagenode',
'meta_keywords' => 'MicroPlex Library Preparation Kit',
'meta_description' => 'MicroPlex Library Preparation Kits v3 have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg.',
'modified' => '2023-04-06 12:08:35',
'created' => '2019-07-16 16:03:58',
'ProductsRelated' => array(
[maximum depth reached]
),
'Image' => array(
[maximum depth reached]
)
)
),
'Application' => array(
(int) 0 => array(
'id' => '14',
'position' => '10',
'parent_id' => '3',
'name' => 'DNA/RNA library preparation',
'description' => '<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p><span style="font-weight: 400;">Most of the major next-generation sequencing platforms require ligation of specific adaptor oligos to </span><a href="../applications/dna-rna-shearing"><span style="font-weight: 400;">fragmented DNA or RNA</span></a><span style="font-weight: 400;"> prior to sequencing</span></p>
<p><span style="font-weight: 400;">After input DNA has been fragmented, it is end-repaired and blunt-ended</span><span style="font-weight: 400;">. The next step is a A-tailing in which dAMP is added to the 3´ end of the blunt phosphorylated DNA fragments to prevent concatemerization and to allow the ligation of adaptors with complementary dT overhangs. In addition, barcoded adapters can be incorporated to facilitate multiplexing prior to or during amplification.</span></p>
<center><img src="https://www.diagenode.com/img/categories/library-prep/flux.png" /></center>
<p><span style="font-weight: 400;">Diagenode offers a comprehensive product portfolio for library preparation:<br /></span></p>
<strong><a href="https://www.diagenode.com/en/categories/Library-preparation-for-RNA-seq">D-Plex RNA-seq Library Preparation Kits</a></strong><br />
<p><span style="font-weight: 400;">Diagenode’s new RNA-sequencing solutions utilize the innovative c</span><span style="font-weight: 400;">apture and a</span><span style="font-weight: 400;">mplification by t</span><span style="font-weight: 400;">ailing and s</span><span style="font-weight: 400;">witching”</span><span style="font-weight: 400;">, a ligation-free method to produce DNA libraries for next generation sequencing from low input amounts of RNA. </span><span style="font-weight: 400;"></span><a href="../categories/Library-preparation-for-RNA-seq">Learn more</a></p>
<strong><a href="../categories/library-preparation-for-ChIP-seq">ChIP-seq and DNA sequencing library preparation solutions</a></strong><br />
<p><span style="font-weight: 400;">Our kits have been optimized for DNA library preparation used for next generation sequencing for a wide range of inputs. Using a simple three-step protocols, our</span><a href="http://www.diagenode.com/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns"><span style="font-weight: 400;"> </span></a><span style="font-weight: 400;">kits are an optimal choice for library preparation from DNA inputs down to 50 pg. </span><a href="../categories/library-preparation-for-ChIP-seq">Learn more</a></p>
<a href="../p/bioruptor-pico-sonication-device"><span style="font-weight: 400;"></span><strong>Bioruptor Pico - short fragments</strong></a><a href="../categories/library-preparation-for-ChIP-seq-and-DNA-sequencing"><span style="font-weight: 400;"></span></a><br />
<p><span style="font-weight: 400;"></span><span style="font-weight: 400;">Our well-cited Bioruptor Pico is the shearing device of choice for chromatin and DNA fragmentation. Obtain uniform and tight fragment distributions between 150bp -2kb. </span><a href="../p/bioruptor-pico-sonication-device">Learn more</a></p>
<strong><a href="../p/megaruptor2-1-unit"><span href="../p/bioruptor-pico-sonication-device">Megaruptor</span>® - long fragments</a></strong><a href="../p/bioruptor-pico-sonication-device"><span style="font-weight: 400;"></span></a><a href="../categories/library-preparation-for-ChIP-seq-and-DNA-sequencing"><span style="font-weight: 400;"></span></a><br />
<p><span style="font-weight: 400;"></span><span style="font-weight: 400;">The Megaruptor is designed to shear DNA from 3kb-75kb for long-read sequencing. <a href="../p/megaruptor2-1-unit">Learn more</a></span></p>
<span href="../p/bioruptor-pico-sonication-device"></span><span style="font-weight: 400;"></span></div>
</div>',
'in_footer' => false,
'in_menu' => true,
'online' => true,
'tabular' => true,
'slug' => 'library-preparation',
'meta_keywords' => 'Library preparation,Next Generation Sequencing,DNA fragments,MicroPlex Library,iDeal Library',
'meta_description' => 'Diagenode offers a comprehensive product portfolio for library preparation such as CATS RNA-seq Library Preparation Kits,ChIP-seq and DNA sequencing library preparation solutions',
'meta_title' => 'Library preparation for Next Generation Sequencing (NGS) | Diagenode',
'modified' => '2021-03-10 03:27:16',
'created' => '2014-08-16 07:47:56',
'ProductsApplication' => array(
[maximum depth reached]
)
)
),
'Category' => array(
(int) 0 => array(
'id' => '124',
'position' => '1',
'parent_id' => '15',
'name' => 'Library preparation for ChIP-seq',
'description' => '<div class="row">
<div class="large-12 columns text-justify">
<p>Library preparation following ChIP can be challenging due to the limited amount of DNA recovered after ChIP. Diagenode has developed the optimal solutions for ChIP-seq using two different approaches: the ligation-based library preparation on purified DNA or the tagmentation-based ChIPmentation.</p>
</div>
</div>
<div class="row extra-spaced">
<div class="large-12 columns"><center><a href="https://www.diagenode.com/en/pages/form-microplex-promo" target="_blank"></a></center></div>
</div>
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div id="portal" class="main-portal">
<div class="portal-inner"><nav class="portal-nav">
<ul data-tab="" class="tips-menu">
<li><a href="#panel1" class="tips portal button">Ligation-based library prep</a></li>
<li><a href="#panel2" class="tips portal button">ChIPmentation</a></li>
<li><a href="#panel3" class="tips portal button">Kit choice guide</a></li>
<li><a href="#panel4" class="tips portal button">Resources</a></li>
<li><a href="#panel5" class="tips portal button">FAQs</a></li>
</ul>
</nav></div>
</div>
<div class="tabs-content">
<div class="content active" id="panel1">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v5" style="color: #13b29c;"><i class="fa fa-caret-right"></i> Standard input library prep</a>
<div id="v5" class="content">
<div class="small-12 medium-12 large-12 columns">
<p>The <strong>iDeal Library Preparation Kit</strong> reliably converts DNA into indexed libraries for next-generation sequencing, with input amounts down to <strong>5 ng</strong>. Our kit offers a simple and fast workflow, high yields, and ready-to-sequence DNA on the Illumina platform.</p>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> <strong>Sample</strong>: Fragmented dsDNA</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Input</strong>: 5 ng – 1 µg</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Fast protocol</strong>: 3 hours</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Easy processing</strong>: 3 steps</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Indexing</strong>: single indexes for multiplexing up to 24 samples</li>
<li><i class="fa fa-arrow-circle-right"></i> Manual and automated protocols available</li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<ul class="square">
<li>MeDIP-seq library prep</li>
<li>Genomic DNA sequencing</li>
<li>High input ChIP-seq</li>
</ul>
</div>
<div class="extra-spaced">
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010020</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" style="color: #b21329;" target="_blank">iDeal Library Preparation Kit x24 (incl. Index Primer Set 1)</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010021</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" style="color: #b21329;" target="_blank">Index Primer Set 2 (iDeal Lib. Prep Kit x24)</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
</li>
</ul>
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v4" style="color: #13b29c;"><i class="fa fa-caret-right"></i> Low input library prep</a>
<div id="v4" class="content active"><center><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" target="_blank"><img src="https://www.diagenode.com/img/banners/banner-microplex-v3-580.jpg" class="extra-spaced" /></a></center>
<div align="center"><a href="https://www.diagenode.com/pages/form-microplex3" class="center alert radius button extra-spaced"><i class="fa fa-info"></i> Contact us</a></div>
<div class="extra-spaced">
<p>Diagenode’s <strong>MicroPlex Library Preparation kits</strong> have been extensively validated for ChIP-seq samples. Generated libraries are compatible with single-end or paired-end sequencing. MicroPlex chemistry (using stem-loop adapters ) is specifically developed and optimized to generate DNA libraries with high molecular complexity from the lowest input amounts. Only <strong>50 pg to 50 ng</strong> of fragmented double-stranded DNA is required for library preparation. The entire <strong>three-step workflow</strong> takes place in a <strong>single tube</strong> or well in about <strong>2 hours</strong>. No intermediate purification steps and no sample transfers are necessary to prevent handling errors and loss of valuable samples.</p>
</div>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> <strong>Sample</strong>: Fragmented dsDNA</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Low input</strong>: 50 pg – 50 ng</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Fast protocol</strong>: 2 hours</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>Easy processing</strong>: 3 steps in 1 tube</li>
<li><i class="fa fa-arrow-circle-right"></i> <strong>No intermediate purification</strong></li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
<li><i class="fa fa-arrow-circle-right"></i> Manual and automated protocols available</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<ul class="square">
<li>ChIP-seq library prep from ChIP-derived DNA</li>
<li>Low input DNA sequencing</li>
</ul>
</div>
<h2>Two versions are available:</h2>
<ul class="accordion" data-accordion="">
<li class="accordion-navigation"><a href="#v2" style="color: #13b29c;"><i class="fa fa-caret-right"></i> MicroPlex Library Preparation Kit v2 with single indexes</a>
<div id="v2" class="content">
<p>The MicroPlex Library Preparation Kit v2 contains all necessary reagents including single indexes for multiplexing up to 48 samples using single barcoding.</p>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010012</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v2 (12 indexes)</a></td>
<td class="format">12 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</li>
<li class="accordion-navigation"><a href="#v3" style="color: #13b29c;"><i class="fa fa-caret-right"></i> MicroPlex Library Preparation Kit v3 with dual indexes <strong><span class="diacol">NEW!</span></strong></a>
<div id="v3" class="content active">
<p>In this version the library preparation reagents and the dual indexes are available separately allowing for the flexibility choosing the number of indexes. MicroPlex v3 has multiplexing capacities up to 384 samples.</p>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010001</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v3 /48 rxns</a></td>
<td class="format">48 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010002</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" style="color: #b21329;" target="_blank">MicroPlex Library Preparation Kit v3 /96 rxns</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
<h4>DUAL INDEXES</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C05010003</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" style="color: #b21329;" target="_blank">24 Dual indexes for MicroPlex Kit v3</a></td>
<td class="format">48 rxns</td>
<td><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010004</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set I</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010005</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set II</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010006</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set III</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C05010007</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" style="color: #b21329;" target="_blank">96 Dual indexes for MicroPlex Kit v3 – Set IV</a></td>
<td class="format">96 rxns</td>
<td><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</li>
</ul>
</div>
</li>
</ul>
</div>
</div>
</div>
<div class="content active" id="panel2">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<p>The TAG Kit for ChIPmentation offers an optimized ChIP-seq library preparation solution based on tagmentation. This kit includes reagents for tagmentation-based library preparation integrated in the IP and is compatible with any ChIP protocol based on magnetic beads. The primer indexes for multiplexing must be purchased separately and are available as a reference: <a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" target="_blank">24 SI for ChIPmentation</a>, Cat. No. C01011031. Alternatively, for histone marks, Diagenode proposes the complete solution (including all buffers for ChIP, tagmentation and multiplexing): <a href="https://www.diagenode.com/en/p/manual-chipmentation-kit-for-histones-24-rxns" target="_blank">ChIPmentation for Histones</a>.</p>
</div>
<div class="extra-spaced">
<h2>Features</h2>
<ul class="nobullet">
<li><i class="fa fa-arrow-circle-right"></i> Sample: chromatin-antibody-magnetic beads complexes</li>
<li><i class="fa fa-arrow-circle-right"></i> Input: chromatin from 5 K – 4 M cells</li>
<li><i class="fa fa-arrow-circle-right"></i> Easy and fast protocol</li>
<li><i class="fa fa-arrow-circle-right"></i> Compatible with any ChIP protocol based on magnetic beads</li>
<li><i class="fa fa-arrow-circle-right"></i> No adapter dimers</li>
<li><i class="fa fa-arrow-circle-right"></i> Sequencing technology: Illumina</li>
</ul>
</div>
<div class="extra-spaced">
<h2>Applications</h2>
<p class="lead"><em><strong>TAG kit for ChIPmentation</strong></em></p>
<ul class="square">
<li>ChIPmentation library preparation</li>
</ul>
<p class="lead"><em><strong>24 SI for for ChIPmentation</strong></em></p>
<ul class="square">
<li>ChIPmentation library preparation</li>
<li>Tagmentation-based library preparation methods like ATAC-seq, CUT&Tag</li>
</ul>
</div>
<h4>KITS</h4>
<table>
<thead>
<tr>
<th>Cat. No.</th>
<th>Product</th>
<th>Format</th>
<th width="120"></th>
</tr>
</thead>
<tbody class="list">
<tr>
<td class="catalog_number"><span class="success label">C01011030</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" style="color: #b21329;" target="_blank">TAG Kit for ChIPmentation</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
<tr>
<td class="catalog_number"><span class="success label">C01011031</span></td>
<td class="name"><a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" style="color: #b21329;" target="_blank">24 SI for ChIPmentation</a></td>
<td class="format">24 rxns</td>
<td><a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" class="tiny details button radius" target="_blank"><i class="fa fa-eye"></i></a></td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
<div class="content" id="panel3">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<h3 class="text-center diacol"><em>How to choose your library preparation kit?</em></h3>
</div>
<table class="noborder">
<tbody>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Sample</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Chromatin-antibody-beads complex</p>
</td>
<td colspan="2">
<p class="text-center" style="font-size: 15px;">Purified DNA</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Purified DNA</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td colspan="2"><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Application</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">ChIPmentation</p>
</td>
<td colspan="2">
<p class="text-center" style="font-size: 15px;">ChIP-seq library prep<br /> Low input DNA sequencing</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">MeDIP-seq library prep<br /> Genomic DNA sequencing<br /> High input ChIP-seq</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td colspan="2"><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Input</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Chromatin: 5 K to 4 M cells</p>
</td>
<td colspan="2"">
<p class="text-center" style="font-size: 15px;">DNA: 50 pg – 50 ng</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">DNA: 5 ng – 1 µg</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow-45-left.png" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow-45-right.png" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Multiplexing</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 24 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 384 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 48 samples</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Up to 24 samples</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Indexes</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Dual indexes (DI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
<td>
<p class="text-center" style="font-size: 15px;">Single indexes (SI)</p>
</td>
</tr>
<tr style="background-color: #fff;">
<td></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
<td><center><img src="https://www.diagenode.com/img/long-arrow.gif" /></center></td>
</tr>
<tr valign="top">
<td class="text-right">
<p style="font-size: 15px;"><strong>Kit</strong></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>TAG Kit for ChIPmentation</strong><br /> (indexes not included in the kit)</p>
<p class="text-center"><strong>Kit</strong><br /> <a href="https://www.diagenode.com/en/p/tag-kit-for-chipmentation-24" target="_blank">C01011030 – 24 rxns</a></p>
<p class="text-center"><strong>Single indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-si-for-chipmentation" target="_blank">C01011031 – 24 SI/24 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>MicroPlex Library Preparation Kit v3</strong><br />(dual indexes not included in the kit)</p>
<p class="text-center"><strong>Kit</strong><br /> <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-48-rxns" target="_blank">C05010001 - 48 rxns</a><br /> <a href="https://www.diagenode.com/en/p/microplex-lib-prep-kit-v3-96-rxns" target="_blank">C05010002 - 96 rxns</a></p>
<br />
<p class="text-center"><strong>Unique dual indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1" target="_blank">C05010008 - Set I 24 UDI / 24 rxns</a><br /> <a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2" target="_blank">C05010009 - Set II 24 UDI/ 24 rxns</a></p>
<p class="text-center"><strong>Dual indexes</strong><br /> <a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns" target="_blank">C05010003 - 24 DI/ 48 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1" target="_blank">C05010004 - Set I 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2" target="_blank">C05010005 - Set II 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3" target="_blank">C05010006 - Set III 96 DI/ 96 rxns</a><br /> <a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4" target="_blank">C05010007 - Set IV 96 DI/ 96 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>MicroPlex Library Preparation Kit v2</strong><br />(single indexes included in the kit)</p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x12-12-indices-12-rxns" target="_blank">C05010012 - 12 SI/ 12 rxns</a><br /> <a href="https://www.diagenode.com/en/p/microplex-library-preparation-kit-v2-x48-12-indices-48-rxns" target="_blank">C05010013 - 12 SI/ 48 rxns</a></p>
</td>
<td>
<p class="text-center" style="font-size: 15px;"><strong>iDeal Library Preparation Kit</strong><br />(Set 1 of indexes included in the kit)</p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/ideal-library-preparation-kit-x24-incl-index-primer-set-1-24-rxns" target="_blank">C05010020 - 12 SI/ 24 rxns</a></p>
<p class="text-center" style="font-size: 15px;"><strong>Index Primer Set 2</strong></p>
<p class="text-center"><a href="https://www.diagenode.com/en/p/ideal-library-index-primer-set-2-24-rxns" target="_blank">C05010021 - 12 SI/ 24 rxns</a></p>
</td>
</tr>
</tbody>
</table>
</div>
</div>
</div>
<div class="content" id="panel4">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p>Combined chromatin immunoprecipitation and next-generation sequencing (ChIP-seq) has become the gold standard to investigate genome-wide epigenetic profiles. However, ChIP from a limited amount of cells has been a challenge. Here we provide a complete and robust workflow solution for successful ChIP-seq from small numbers of cells using the True MicroChIP kit and MicroPlex Library Preparation kit.</p>
<blockquote><span class="label-green" style="margin-bottom: 16px; margin-left: -22px;">APPLICATION NOTE</span>
<div class="row">
<div class="small-12 medium-6 large-6 columns"><center><img src="https://www.diagenode.com/img/categories/microplex/chip-efficiency-on-10000-cells.jpg" /></center>
<p><small><em>ChIP efficiency on 10,000 cells</em></small></p>
</div>
<div class="small-12 medium-6 large-6 columns">
<p><strong>From minuscule amounts to magnificent results:</strong><br /> reliable ChIP-seq data from 10,000 cells with the True MicroChIP™ and the MicroPlex Library Preparation™ kits.</p>
<a href="https://www.diagenode.com/files/application_notes/True_MicroChIP_and_MicroPlex_kits_Application_Note.pdf" class="details small button" target="_blank">DOWNLOAD</a></div>
</div>
</blockquote>
<blockquote><span class="label-green" style="margin-bottom: 16px; margin-left: -22px;">APPLICATION NOTE</span>
<div class="row">
<div class="small-12 medium-6 large-6 columns"><center><img src="https://www.diagenode.com/img/categories/microplex/quality-control-check.jpg" /></center>
<p class="text-left"><small><em>Quality control check of a ChIP-seq library on the Fragment Analyzer. High Efficiency ChIP performed on 10,000 cells</em></small></p>
</div>
<div class="small-12 medium-6 large-6 columns">
<p class="text-left"><strong>Best Workflow Practices for ChIP-seq Analysis with Small Samples</strong></p>
<a href="https://www.diagenode.com/files/application_notes/Diagenode_AATI_Joint.pdf" class="details small button" target="_blank">DOWNLOAD</a></div>
</div>
</blockquote>
</div>
</div>
</div>
<div class="content" id="panel5">
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<div class="extra-spaced">
<h2>TAG Kit for ChIPmentation</h2>
<ol>
<li><strong>What is the difference between tagmentation and ChIPmentation?</strong><br />Tagmentation is a reaction where an enzyme (a transposase) cleaves DNA and incorporates sequencing adaptors at the ends of the fragments in one step. In our ChIPmentation technology we combine chromatin immunoprecipitation and tagmentation in one streamlined workflow where the tagmentation step occurs directly on chromatin.<br /><br /></li>
<li><strong>What is the expected concentration of ChIPmentation libraries?</strong><br />The concentration of libraries that you need to reach will depend on the sensitivity of the machine and kits that you will use to perform the quality control and the sequencing of your libraries. Usually a concentration of 4-8 ng/μl is enough for a quality control using the Qubit High Sensitivity assay (ThermoFischer Scientific) and the High Sensitivity chip for BioAnalyzer (Agilent) and for sequencing on Illumina HiSeq3000/4000.<br /><br /></li>
<li><strong>Does the ChIPmentation approach work on plants?</strong><br />Our ChIPmentation solution has been validated on human cells and we do not have any data on plants. It should be compatible. We would recommend using our Universal Plant ChIP Kit in combination with the TAG Kit for ChIPmentation and the 24 SI for ChIPmentation.<br /><br /></li>
<li><strong>What is the size of the fragments after the tagmentation?</strong><br />The size of the fragments at the end of the ChIPmentation protocol can vary depending on many parameters like the shearing efficiency, the antibody used or the tagmentation time. However, with our standard protocol we usually obtain a library peak which is around 200-300 bp (see example of results at the end of the manual). If many fragments larger than 500 bp are present , the best would be to contact your sequencing provider to ask what their requirements are, because it can vary depending on the sequencer. If you want to remove the large fragments you can use the size selection protocol described in the manual.<br /><br /></li>
<li><strong>What is the size of the adapters?</strong><br />The sum of the adapters is 128 bp.</li>
</ol>
</div>
<div class="extra-spaced">
<h2>MicroPlex Library Preparation Kit</h2>
<ol>
<li><strong>Can I use the available Illumina primers and validate them with the MicroPlex Kit v2?</strong><br /> Although the final flanking sequences of MicroPlex are the same as those used by Illumina, the PCR primers are not identical and part of them is supplied with the buffer. For this reason Illumina primers will not work as substitute.<br /><br /></li>
<li><strong>The BioAnalyzer profile of purified library shows the presence of low molecular weight peaks (primers/adaptors) in the samples. Should I re- purify the samples or they can be used directly to the sequencing? If the second purification is recommended, which ratio sample/AMPure beads should I use?</strong><br /> You can do a second round of purification using 1:1 ratio of AMPure beads to sample and this should get rid of the majority of the dimers.<br /><br /></li>
<li><strong>I am going to use the MicroPlex Library Preparation Kit v2 on ChIP samples . Our thermocycler has ramp rate 1.5°/s max while the protocol recommends using a ramp rate 3 to 5°/s. How would this affect the library prep?</strong><br /> We have not used a thermocycler with a ramp rate of 1.5 °C, which seems faster than most of thermocyclers. Too fast of a ramp rate may affect the primer annealing and ligation steps.<br /><br /></li>
<li><strong>What is the function of the replication stop site in the adapter loops?</strong><br /> The replication stop site in the adaptor loops function to stop the polymerase from continuing to copy the rest of the stem loop.<br /><br /></li>
<li><strong>I want to do ChIP-seq. Which ChIP-seq kit can I use for sample preparation prior to Microplex Library Preparation Kit v2?</strong><br /> In our portfolio there are several ChIP-seq kits compatible with Microplex Library Preparation Kit v2. Depending on your sample type and target studied you can use the following kits: iDeal ChIP-seq Kit for Transcription Factors (Cat. No. C01010055), iDeal ChIP-seq Kit for Histones (Cat. No. C01010051), True MicroChIP kit (Cat. No. C01010130), Universal Plant ChIP-seq Kit (Cat. No. C01010152). All these kits exist in manual and automated versions.<br /><br /></li>
<li><strong>Is Microplex Library Preparation Kit v2 compatible with exome enrichment methods?</strong><br /> Microplex Library Preparation Kit v2 is compatible with major exome and target enrichment products, including Agilent SureSelect<sup>®</sup>, Roche NimbleGen<sup>®</sup> SeqCap<sup>®</sup> EZ and custom panels.<br /><br /></li>
<li><strong>What is the nick that is mentioned in the kit method overview?</strong><br /> The nick is simply a gap between a stem adaptor and 3’ DNA end, as shown on the schema in the kit method overview.<br /><br /></li>
<li><strong>Are the indexes of the MicroPlex library preparation kit v2 located at i5 or i7?</strong><br /> The libraries generated with the MicroPlex kit v2 contain indices located at i7.<br /><br /></li>
<li><strong>Is there a need to use custom index read primers for the sequencing to read the 8nt iPCRtags?</strong><br /> There is no need for using custom Sequencing primer to sequence MicroPlex libraires. MicroPlex libraries can be sequenced using standard Illumina Sequencing kits and protocols.<br /><br /></li>
<li><strong>What is the advantage of using stem-loop adapter in the MicroPlex kit?</strong><br /> There are several advantages of using stem-loop adaptors. First of all, stem-loop adaptors prevent from self-ligation thus increases the ligation efficiency between the adapter and DNA fragment. Moreover, the background is reduced using ds adaptors with no single-stranded tails. Finally, adaptor-adaptor ligation is reduced using blocked 5’ ends.<br /><br /></li>
</ol>
</div>
<div class="extra-spaced">
<h2>IDeal Library Preparation Kit</h2>
<ol>
<li><strong>Are the index from the iDeal library Prep kit compatible with the MicroPlex library prep kit?</strong><br /> No, it is important to use only the indexes provided in the MicroPlex kit to ensure proper library preparation with this kit</li>
</ol>
</div>
</div>
</div>
</div>
</div>
</div>
</div>
<script>// <![CDATA[
$(document).ready(function() {
$('ul.tips-menu li a:first').trigger('focus');
});
// ]]></script>',
'no_promo' => false,
'in_menu' => true,
'online' => true,
'tabular' => true,
'hide' => true,
'all_format' => false,
'is_antibody' => false,
'slug' => 'library-preparation-for-ChIP-seq',
'cookies_tag_id' => null,
'meta_keywords' => 'Library preparation,Next Generation Sequencing,DNA fragments,MicroPlex Library,iDeal Library',
'meta_description' => 'Diagenode Kits have been Optimized for DNA library Preparation used for Next Generation Sequencing for a range of inputs.',
'meta_title' => 'DNA Library Preparation Kits,Microplex library Preparation Kits | Diagenode',
'modified' => '2021-12-20 14:30:28',
'created' => '2016-10-21 02:39:19',
'ProductsCategory' => array(
[maximum depth reached]
),
'CookiesTag' => array([maximum depth reached])
)
),
'Document' => array(
(int) 0 => array(
'id' => '1056',
'name' => 'Primer indexes for MicroPlex kit v3',
'description' => '<p>High Performance Library Preparation for Illumina<sup>®</sup> NGS Platforms</p>',
'image_id' => null,
'type' => 'Manual',
'url' => 'files/products/kits/Primer-indexes-for-Microplex-v3.pdf',
'slug' => 'primer-indexes-for-microplex-v3-manual',
'meta_keywords' => '',
'meta_description' => '',
'modified' => '2020-10-22 12:03:45',
'created' => '2019-07-18 11:53:57',
'ProductsDocument' => array(
[maximum depth reached]
)
)
),
'Feature' => array(),
'Image' => array(
(int) 0 => array(
'id' => '1759',
'name' => 'product/kits/chip-kit-icon.png',
'alt' => 'chip kit icon',
'modified' => '2018-01-18 15:26:13',
'created' => '2018-01-18 15:08:38',
'ProductsImage' => array(
[maximum depth reached]
)
)
),
'Promotion' => array(),
'Protocol' => array(),
'Publication' => array(),
'Testimonial' => array(),
'Area' => array(),
'SafetySheet' => array()
)
$country = 'US'
$countries_allowed = array(
(int) 0 => 'CA',
(int) 1 => 'US',
(int) 2 => 'IE',
(int) 3 => 'GB',
(int) 4 => 'DK',
(int) 5 => 'NO',
(int) 6 => 'SE',
(int) 7 => 'FI',
(int) 8 => 'NL',
(int) 9 => 'BE',
(int) 10 => 'LU',
(int) 11 => 'FR',
(int) 12 => 'DE',
(int) 13 => 'CH',
(int) 14 => 'AT',
(int) 15 => 'ES',
(int) 16 => 'IT',
(int) 17 => 'PT'
)
$outsource = false
$other_formats = array()
$edit = ''
$testimonials = ''
$featured_testimonials = ''
$related_products = '<li>
<div class="row">
<div class="small-12 columns">
<a href="/en/p/microplex-lib-prep-kit-v3-48-rxns"><img src="/img/product/kits/chip-kit-icon.png" alt="chip kit icon" class="th"/></a> </div>
<div class="small-12 columns">
<div class="small-6 columns" style="padding-left:0px;padding-right:0px;margin-top:-6px;margin-left:-1px">
<span class="success label" style="">C05010001</span>
</div>
<div class="small-6 columns text-right" style="padding-left:0px;padding-right:0px;margin-top:-6px">
<!--a href="#" style="color:#B21329"><i class="fa fa-info-circle"></i></a-->
<!-- BEGIN: ADD TO CART MODAL --><div id="cartModal-3032" class="reveal-modal small" data-reveal aria-labelledby="modalTitle" aria-hidden="true" role="dialog">
<form action="/en/carts/add/3032" id="CartAdd/3032Form" method="post" accept-charset="utf-8"><div style="display:none;"><input type="hidden" name="_method" value="POST"/></div><input type="hidden" name="data[Cart][product_id]" value="3032" id="CartProductId"/>
<div class="row">
<div class="small-12 medium-12 large-12 columns">
<p>Add <input name="data[Cart][quantity]" placeholder="1" value="1" min="1" style="width:60px;display:inline" type="number" id="CartQuantity" required="required"/> <strong> MicroPlex Library Preparation Kit v3 /48 rxns</strong> to my shopping cart.</p>
<div class="row">
<div class="small-6 medium-6 large-6 columns">
<button class="alert small button expand" onclick="$(this).addToCart('MicroPlex Library Preparation Kit v3 /48 rxns',
'C05010001',
'1555',
$('#CartQuantity').val());" name="checkout" id="checkout" value="checkout" type="submit">Checkout</button> </div>
<div class="small-6 medium-6 large-6 columns">
<button class="alert small button expand" onclick="$(this).addToCart('MicroPlex Library Preparation Kit v3 /48 rxns',
'C05010001',
'1555',
$('#CartQuantity').val());" name="keepshop" id="keepshop" type="submit">Keep shopping</button> </div>
</div>
</div>
</div>
</form><a class="close-reveal-modal" aria-label="Close">×</a></div><!-- END: ADD TO CART MODAL --><a href="#" id="microplex-lib-prep-kit-v3-48-rxns" data-reveal-id="cartModal-3032" class="" style="color:#B21329"><i class="fa fa-cart-plus"></i></a>
</div>
</div>
<div class="small-12 columns" >
<h6 style="height:60px">MicroPlex Library Preparation Kit v3 /48 rxns</h6>
</div>
</div>
</li>
'
$related = array(
'id' => '3032',
'antibody_id' => null,
'name' => 'MicroPlex Library Preparation Kit v3 /48 rxns',
'description' => '<p><a href="https://www.diagenode.com/files/products/kits/Microplex-library-prep-v3.pdf"><img src="https://www.diagenode.com/img/buttons/bt-manual.png" /></a></p>
<p>Diagenode’s <strong>MicroPlex Library Preparation Kits v3</strong> have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg. The kit MicroPlex v3 includes all buffers and enzymes necessary for the library preparation. For flexibility of the choice different formats of compatible primer indexes are available separately:</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-dual-indexes-for-microplex-kit-v3-48-rxns">C05010003 - 24 Dual indexes for MicroPlex Kit v3 /48 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-1">C05010004 - 96 Dual indexes for MicroPlex Kit v3 – Set I /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-2">C05010005 - 96 Dual indexes for MicroPlex Kit v3 – Set II /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-3">C05010006 - 96 Dual indexes for MicroPlex Kit v3 – Set III /96 rxns</a></li>
<li><a href="https://www.diagenode.com/en/p/96-dual-indexes-for-microplex-kit-v3-set-4">C05010007 - 96 Dual indexes for MicroPlex Kit v3 – Set IV /96 rxns</a></li>
</ul>
<p style="padding-left: 30px;">NEW! Unique dual indexes :</p>
<ul>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set1">C05010008 - 24 UDI for MicroPlex Kit v3 - Set I</a></li>
<li><a href="https://www.diagenode.com/en/p/24-unique-dual-indexes-for-microplex-kit-v3-set2">C05010009 - 24 UDI for MicroPlex Kit v3 - Set II</a></li>
</ul>
<p>Read more about <a href="https://www.diagenode.com/en/categories/library-preparation-for-ChIP-seq">library preparation for ChIP-seq</a></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label1' => 'Characteristics',
'info1' => '<ul>
<li><strong>1 tube</strong>, <strong>2 hours</strong>, <strong>3 steps</strong> protocol</li>
<li><strong>Input</strong>: 50 pg – 50 ng</li>
<li><strong>Reduce potential bias</strong> - few PCR amplification cycles needed</li>
<li><strong>High sensitivity ChIP-seq</strong> - low PCR duplication rate</li>
<li><strong>Great multiplexing flexibility</strong> with 24 dual indexes (8 nt)</li>
<li><strong>Validated with the IP-<a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit">Star<sup>®</sup></a></strong><a href="https://www.diagenode.com/en/p/sx-8g-ip-star-compact-automated-system-1-unit"> Automated Platform</a></li>
</ul>
<h3>How it works</h3>
<center><img alt="MicroPlex Library Preparation Kit v3 /48 rxns" src="https://www.diagenode.com/img/product/kits/microplex-3-method-overview.png" /></center>
<p style="margin-bottom: 0;"><small><strong>Microplex workflow - protocol with dual indexes</strong><br />An input of 50 pg to 50 ng of fragmented dsDNA is converted into sequencing-ready libraries for Illumina® NGS platforms using a fast and simple 3-step protocol</small></p>
<ul class="accordion" data-accordion="" id="readmore" style="margin-left: 0;">
<li class="accordion-navigation"><a href="#first" style="background: #ffffff; padding: 0rem; margin: 0rem; color: #13b2a2;"><small>Read more about MicroPlex workflow</small></a>
<div id="first" class="content">
<p><small><strong>Step 1. Template Preparation</strong> provides efficient repair of the fragmented double-stranded DNA input.</small></p>
<p><small>In this step, the DNA is repaired and yields molecules with blunt ends.</small></p>
<p><small><strong>Step 2. Library Synthesis.</strong> enables ligation of MicroPlex patented stem- loop adapters.</small></p>
<p><small>In the next step, stem-loop adaptors with blocked 5’ ends are ligated with high efficiency to the 5’ end of the genomic DNA, leaving a nick at the 3’ end. The adaptors cannot ligate to each other and do not have single- strand tails, both of which contribute to non-specific background found with many other NGS preparations.</small></p>
<p><small><strong>Step 3. Library Amplification</strong> enables extension of the template, cleavage of the stem-loop adaptors, and amplification of the library. Illumina- compatible indexes are also introduced using a high-fidelity, highly- processive, low-bias DNA polymerase.</small></p>
<p><small>In the final step, the 3’ ends of the genomic DNA are extended to complete library synthesis and Illumina-compatible indexes are added through a high-fidelity amplification. Any remaining free adaptors are destroyed. Hands-on time and the risk of contamination are minimized by using a single tube and eliminating intermediate purifications.</small></p>
<p><small>Obtained libraries are purified, quantified and sized. The libraries pooling can be performed as well before sequencing.</small></p>
</div>
</li>
</ul>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label2' => '',
'info2' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'label3' => '',
'info3' => '<p></p>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script async="" src="https://edge.fullstory.com/s/fs.js" crossorigin="anonymous"></script>
<script src="chrome-extension://hhojmcideegachlhfgfdhailpfhgknjm/web_accessible_resources/index.js"></script>',
'format' => '48 rxns',
'catalog_number' => 'C05010001',
'old_catalog_number' => '',
'sf_code' => 'C05010001-',
'type' => 'FRE',
'search_order' => '04-undefined',
'price_EUR' => '1750',
'price_USD' => '1555',
'price_GBP' => '1555',
'price_JPY' => '286740',
'price_CNY' => '',
'price_AUD' => '3888',
'country' => 'ALL',
'except_countries' => 'None',
'quote' => false,
'in_stock' => false,
'featured' => true,
'no_promo' => false,
'online' => true,
'master' => true,
'last_datasheet_update' => '',
'slug' => 'microplex-lib-prep-kit-v3-48-rxns',
'meta_title' => 'MicroPlex Library Preparation Kit v3 /48 rxns | Diagenode',
'meta_keywords' => 'MicroPlex Library Preparation Kit',
'meta_description' => 'MicroPlex Library Preparation Kits v3 have been extensively validated for ChIP-seq samples and are optimized to generate DNA libraries with high molecular complexity from the lowest input amounts – down to 50 pg.',
'modified' => '2023-04-06 12:08:35',
'created' => '2019-07-16 16:03:58',
'ProductsRelated' => array(
'id' => '4541',
'product_id' => '3027',
'related_id' => '3032'
),
'Image' => array(
(int) 0 => array(
'id' => '1759',
'name' => 'product/kits/chip-kit-icon.png',
'alt' => 'chip kit icon',
'modified' => '2018-01-18 15:26:13',
'created' => '2018-01-18 15:08:38',
'ProductsImage' => array(
[maximum depth reached]
)
)
)
)
$rrbs_service = array(
(int) 0 => (int) 1894,
(int) 1 => (int) 1895
)
$chipseq_service = array(
(int) 0 => (int) 2683,
(int) 1 => (int) 1835,
(int) 2 => (int) 1836,
(int) 3 => (int) 2684,
(int) 4 => (int) 1838,
(int) 5 => (int) 1839,
(int) 6 => (int) 1856
)
$labelize = object(Closure) {
}
$old_catalog_number = ''
$country_code = 'US'
$label = '<img src="/img/banners/banner-customizer-back.png" alt=""/>'
$document = array(
'id' => '1056',
'name' => 'Primer indexes for MicroPlex kit v3',
'description' => '<p>High Performance Library Preparation for Illumina<sup>®</sup> NGS Platforms</p>',
'image_id' => null,
'type' => 'Manual',
'url' => 'files/products/kits/Primer-indexes-for-Microplex-v3.pdf',
'slug' => 'primer-indexes-for-microplex-v3-manual',
'meta_keywords' => '',
'meta_description' => '',
'modified' => '2020-10-22 12:03:45',
'created' => '2019-07-18 11:53:57',
'ProductsDocument' => array(
'id' => '2775',
'product_id' => '3027',
'document_id' => '1056'
)
)include - APP/View/Products/view.ctp, line 755
View::_evaluate() - CORE/Cake/View/View.php, line 971
View::_render() - CORE/Cake/View/View.php, line 933
View::render() - CORE/Cake/View/View.php, line 473
Controller::render() - CORE/Cake/Controller/Controller.php, line 963
ProductsController::slug() - APP/Controller/ProductsController.php, line 1052
ReflectionMethod::invokeArgs() - [internal], line ??
Controller::invokeAction() - CORE/Cake/Controller/Controller.php, line 491
Dispatcher::_invoke() - CORE/Cake/Routing/Dispatcher.php, line 193
Dispatcher::dispatch() - CORE/Cake/Routing/Dispatcher.php, line 167
[main] - APP/webroot/index.php, line 118
×