Histones are the main constituents of the protein part of chromosomes of eukaryotic cells. They are rich in the amino acids arginine and lysine and have been greatly conserved during evolution. Histones pack the DNA into tight masses of chromatin. Two core histones of each class H2A, H2B, H3 and H4 assemble and are wrapped by 146 base pairs of DNA to form one octameric nucleosome. Histone tails undergo numerous post-translational modifications, which either directly or indirectly alter chromatin structure to facilitate transcriptional activation or repression or other nuclear processes. In addition to the genetic code, combinations of the different histone modifications reveal the so-called “histone code”. Histone methylation and demethylation is dynamically regulated by respectively histone methyl transferases and histone demethylases. Trimethylation of histone H3K9 is associated with inactive genomic regions, satellite repeats and ZNF gene repeats.
H3K9me3 polyclonal antibody
(pAb-193-050)
お見積り
Request a quote for a bulk order for H3K9me3 polyclonal antibody . Please fill out the form here below. Your local sales account manager will get in touch with you shortly and send you a quotation based on your requirements.
Bulk order
| Lot | A2814P |
|---|---|
| Concentration | 2.1 µg/µl |
| Species reactivity | Human, mouse, zebrafish, trout, Daphnia, Arabidopsis, Drosophila, Silene latifolia: positive. Other species: not tested. |
| Type | Polyclonal, ChIP grade, ChIP-seq grade |
| Purity | Affinity purified polyclonal antibody. |
| Host | Rabbit |
| Storage Conditions | Store at -20°C; for long storage, store at -80°C. Avoid multiple freeze-thaw cycles. |
| Storage Buffer | PBS containing 0.05% azide. |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Applications | Suggested dilution | References |
|---|---|---|
| ChIP/ChIP-seq * | 0.5 - 1 μg/ChIP | Fig 1, 2 |
| ELISA | 1:1,000 - 1:10,000 | Fig 3 |
| Dot Blotting | 1:2,000 | Fig 4 |
| Western Blotting | 1:2,000 | Fig 5 |
* Please note that the optimal antibody amount per IP should be determined by the end-user. We recommend testing 0.5-5 µg per IP.
- Validation Data
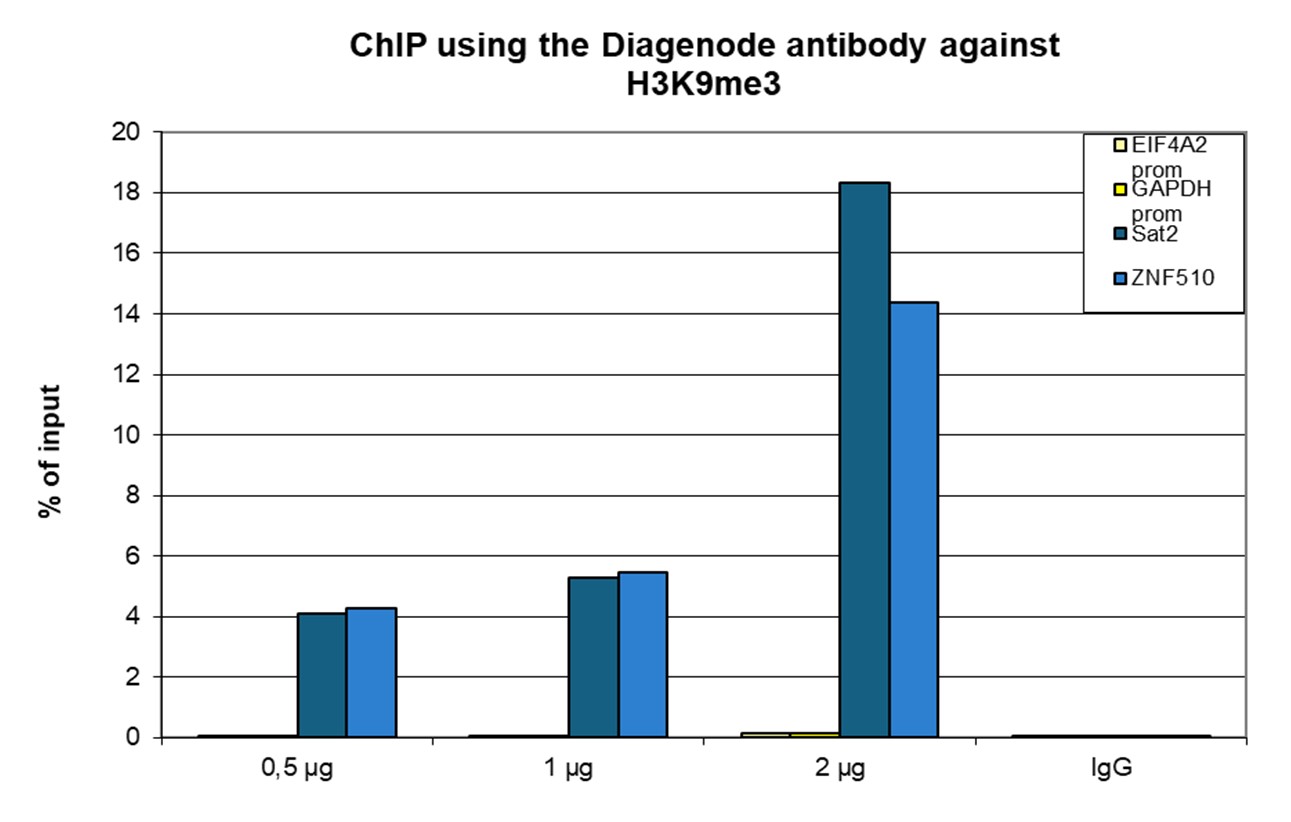
Figure 1. ChIP results obtained with the antibody directed against H3K9me3
ChIP was performed with the antibody against H3K9me3 (cat. no. C15410193) on sheared chromatin from 500,000 K562 cells using the iDeal ChIP-seq kit (cat. no. C01010051). A titration of the antibody consisting of 0.5, 1, and 2 µg per ChIP experiment was analyzed. IgG (1 µg/IP) was used as a negative IP control. Quantitative PCR was performed with primers for the ZNF510 gene and the Sat2 satellite repeat, used as positive controls, and for the EIF4A2 and GAPDH promoters, used as negative controls. The graph shows the recovery, expressed as a % of input (the relative amount of immunoprecipitated DNA compared to input DNA after qPCR analysis).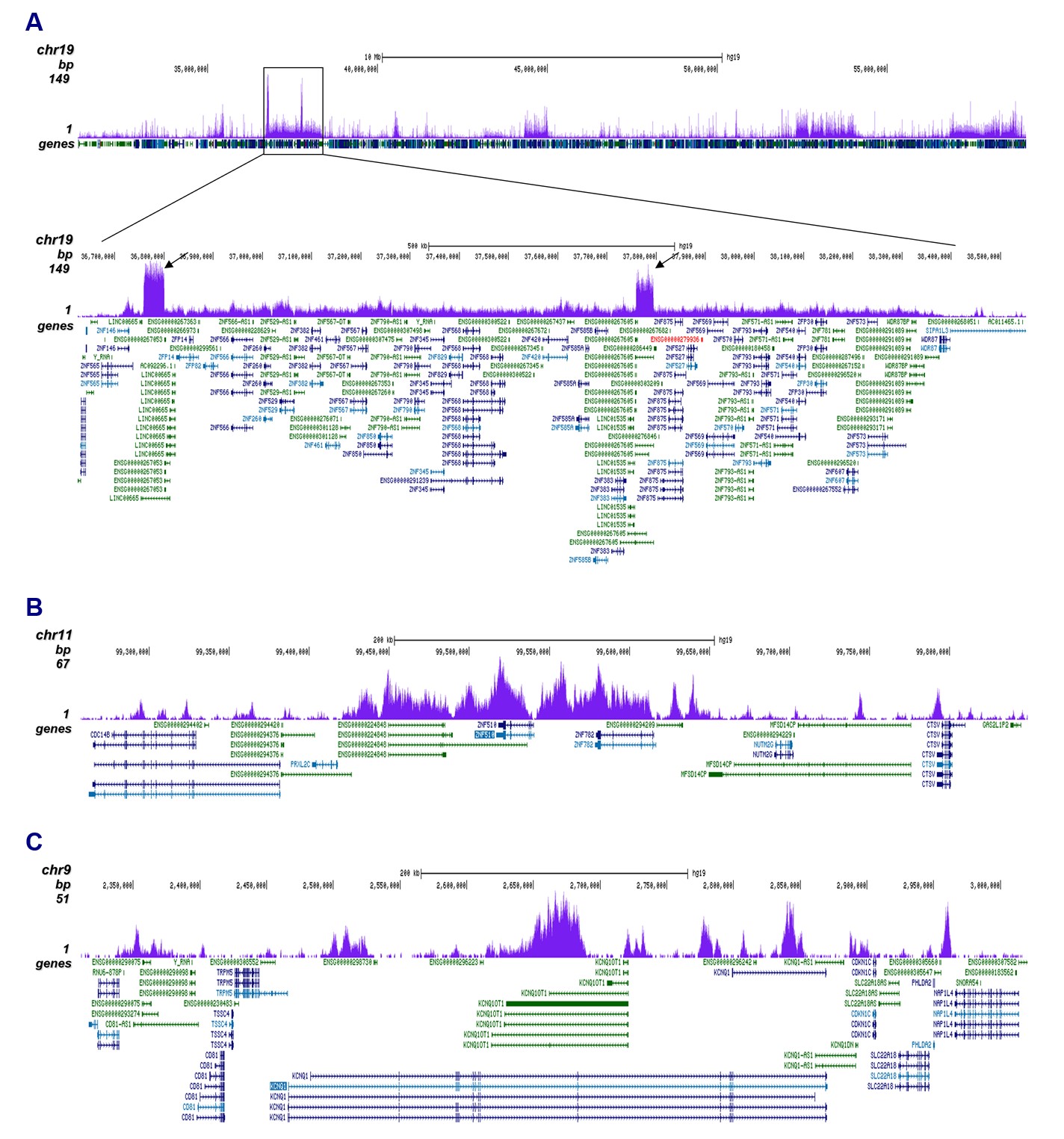
Figure 2. ChIP-seq results obtained with the antibody directed against H3K9me3
ChIP was performed with 1 µg of the antibody against H3K9me3 (cat. no. C15410193) on sheared chromatin from 500,000 K562 cells using the iDeal ChIP-seq kit as described above. The IP'd DNA was subsequently analyzed on an Illumina HiSeq4000. Library preparation, cluster generation, and sequencing were performed according to the manufacturer's instructions. The 50 bp tags were aligned to the human genome using the BWA algorithm. Figure 2A shows the signal distribution along the long arm of chromosome 19 and a zoom-in to an enriched region containing several ZNF repeat genes. The arrows indicate two satellite repeat regions which exhibit a stronger signal. Figures 2B and 2C show the enrichment along the ZNF510 positive control target and at the KCNQ1 imprinted gene.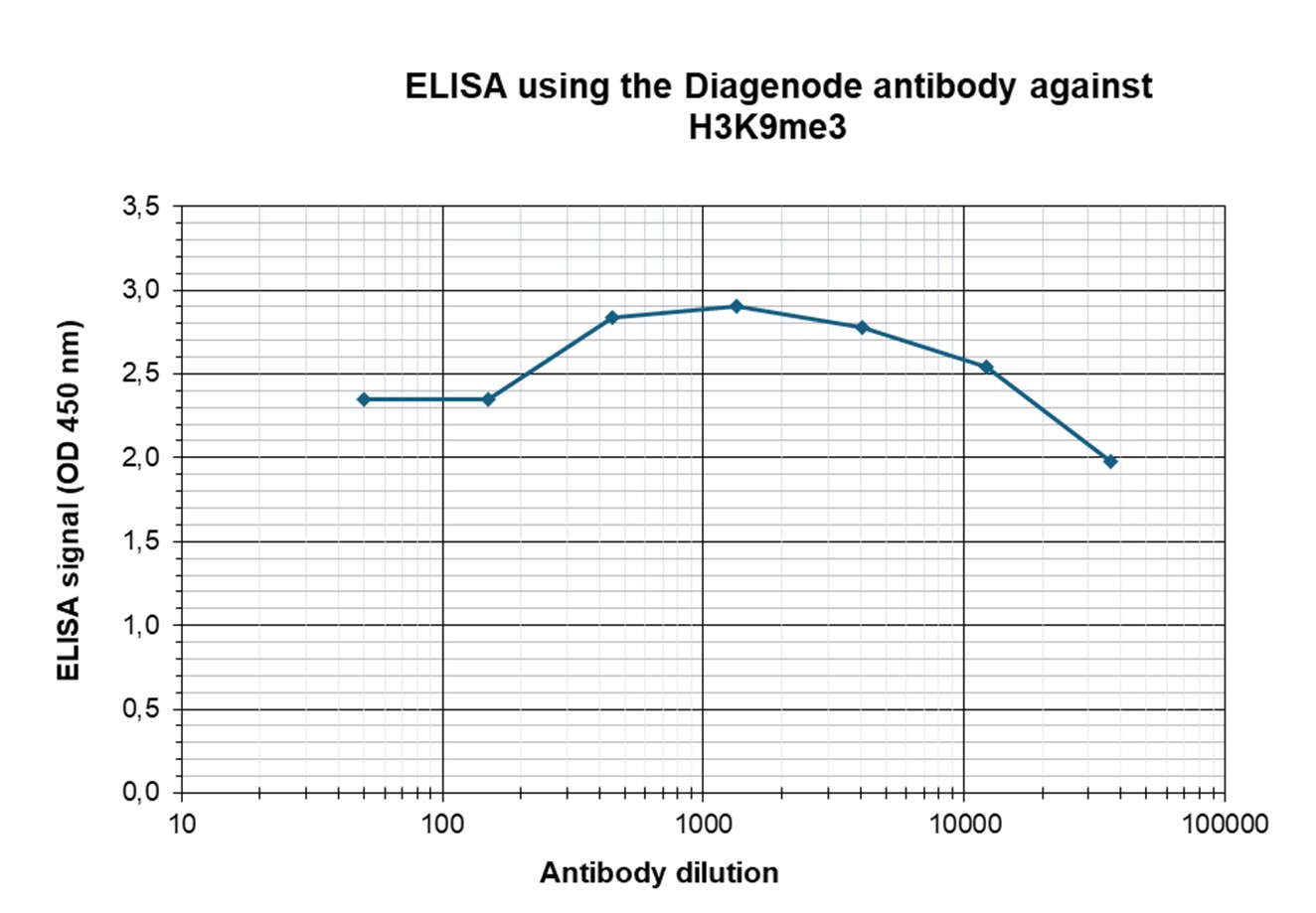
Figure 3. Determination of the antibody titer
To determine the titer of the antibody, an ELISA was performed using a serial dilution of the antibody directed against human H3K9me3 (cat. no. C15410193) in antigen-coated wells. The antigen used was a peptide containing the histone modification of interest. By plotting the absorbance against the antibody dilution (Figure 3), the titer of the antibody was estimated to be 1:183,000.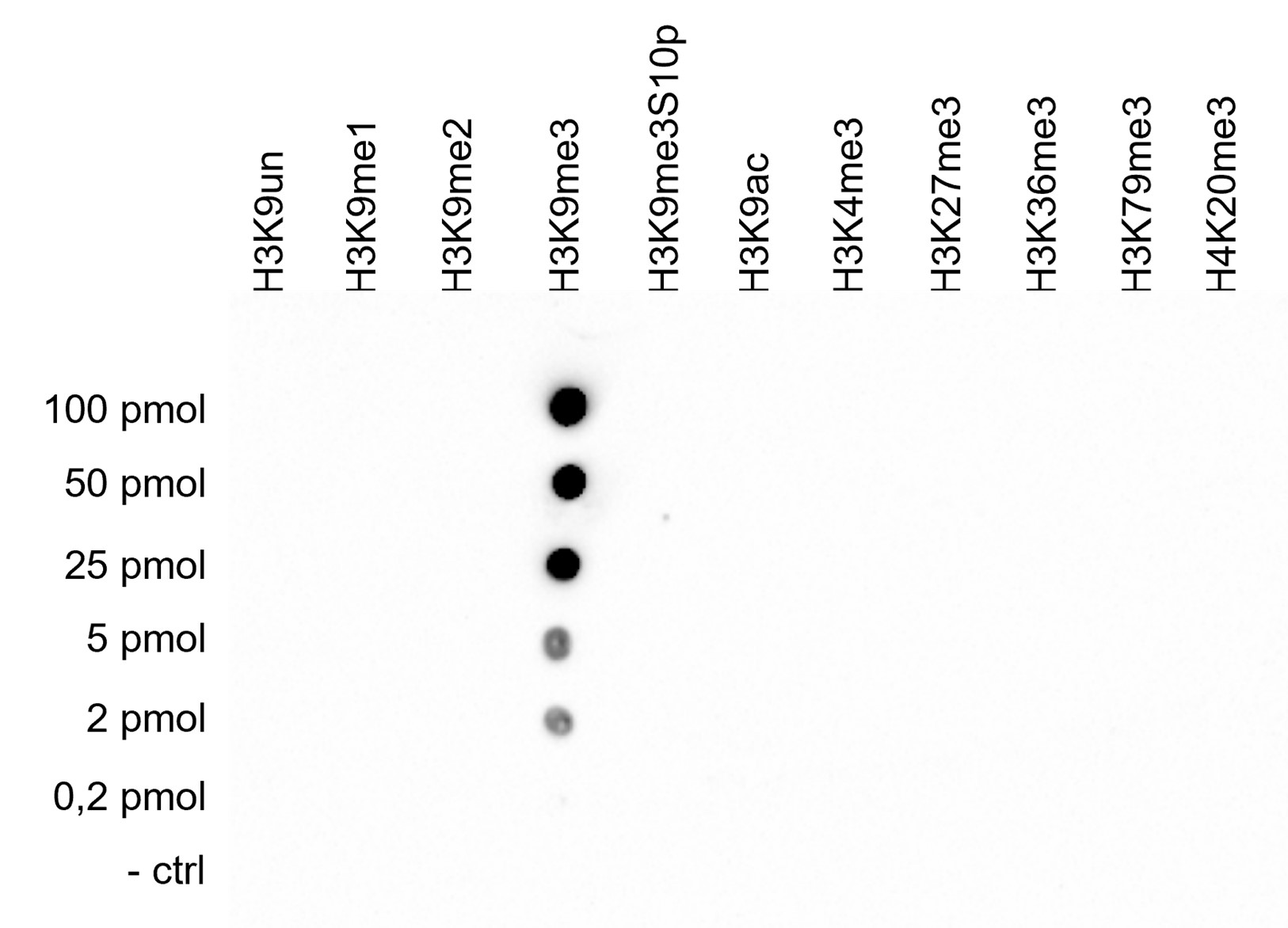
Figure 4. Cross-reactivity tests using the antibody directed against H3K9me3
A Dot Blot analysis was performed to test the cross-reactivity of the antibody against H3K9me3 (cat. no. C15410193) with peptides containing other modifications of histone H3 and H4 and the unmodified sequence of histone H3. One hundred to 0.2 pmol of the peptide containing the respective histone modification were spotted on a membrane. The antibody was used at a dilution of 1:2,000. Figure 4 shows a high specificity of the antibody for the modification of interest.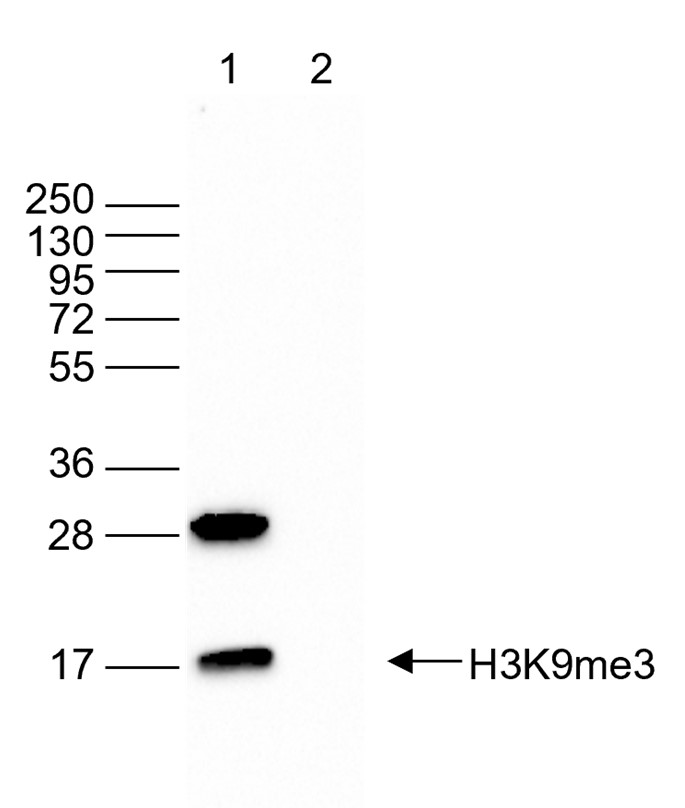
Figure 5. Western blot analysis using the antibody directed against H3K9me3
Western blot was performed on 40 µg of whole cell extracts from HeLa cells (lane 1) and on 1 µg of recombinant histone H3 (lane 2) using the antibody against H3K9me3 (cat. no. C15410193). The antibody was diluted 1:2,000 in TBS-Tween containing 5% skimmed milk. The position of the protein of interest is indicated on the right; the marker (in kDa) is shown on the left. - Target Description
- お客様の声
In life sciences, epigenetics is nowadays the most rapid developing field with new astonishing discoveries made every day. To keep pace with this field, we are in need of reliable tools to foster our research - tools Diagenode provides us with. From antibodies to automated solutions - all from one source and with robust support. Antibodies used in our lab: H3K27me3 polyclonal antibody – Premium, H3K4me3 polyclonal antibody – Premium, H3K9me3 polyclonal antibody – Premium, H3K4me1 polyclonal antibody – Premium, CTCF polyclonal antibody – Classic, Rabbit IgG.
Dr. Florian Uhle, Dept. of Anesthesiology, Heidelberg University Hospital, Germany - 実験手法
ELISA
Enzyme-linked immunosorbent assay. Read moreDB
Dot blotting Read moreWB
Western blot : The quality of antibodies used in this technique is crucial for correct and specific protein identification. Diagenode offers huge selection of highly sensitive and specific western blot-validated antibodies. Learn more about: Load... Read moreChIP-seq (ab)
Read moreChIP-qPCR (ab)
Read more - 資料
Datasheet H3K9me3 pAb-193-050 DATASHEET
Polyclonal antibody raised in rabbit against the region of histone H3 containing the trimethylate...Download Epigenetic Antibodies Brochure BROCHURE
More than in any other immuoprecipitation assays, quality antibodies are critical tools in many e...Download True MicroChIP and MicroPlex kits APPLICATION NOTE
From minuscule amounts to magnificent results: reliable ChIP-seq data from 10,000 cells with the ...Download ChIP kit results with True MicroChIP kit POSTER
Chromatin immunoprecipitation coupled with high-throughput sequencing (ChIP-seq) has become the g...Download Antibodies you can trust POSTER
Epigenetic research tools have evolved over time from endpoint PCR to qPCR to the analyses of lar...Download - Safety sheets
- 出版物

How to properly cite our product/service in your work
We strongly recommend using this: H3K9me3 polyclonal antibody (Hologic Diagenode Cat# C15410193 Lot# A2814P). Click here to copy to clipboard.
Using our products or services in your publication? Let us know!
Cell Cycle Control of Nuclear Metabolism Couples Phosphatidylinositol Signaling to Histone Methylation
Gañez-Zapater, Antoni et al.
Progression through the cell cycle requires coordinated regulation of transcription, chromatin state, and cellular metabolism. While metabolic enzymes are known to localize the nucleus and influence chromatin states, how nuclear metabolism itself oscillates during the cell cycle remains unexplored. Here, we combin...Stage-specific epigenetic priming amplifies gene activation during lineage commitment
Foertsch, Arion et al.
Neural progenitor cells exhibit developmental plasticity as they can commit to distinct developmental trajectories. The male-specific lethal complex (MSLc) is linked to multiple developmental disorders, suggesting a role in neural fate commitment. To dissect MSLc function, we used a multipronged approach combini...Coordinated control of genome-nuclear lamina interactions by topoisomerase 2B and lamin B receptor
van Schaik, Tom et al.
Lamina-associated domains (LADs) are megabase-sized genomic regions anchored to the nuclear lamina (NL). Factors controlling the interactions of the genome with the NL have largely remained elusive. Here, we identified DNA topoisomerase 2 beta (TOP2B) as a regulator of these interactions. TOP2B binds predominant...The senolytic cocktail, dasatinib and quercetin, impacts the chromatin structure of both young and senescent vascular smooth muscle cells
Agnieszka Gadecka et al.
One promising strategy to alleviate aging symptoms is the treatment with senolytics that is compounds which selectively eliminate senescent cells. Some therapies aim to reduce symptoms of cellular senescence without senescent cell eradication (senomorphic activity). However, senotherapies raise many questions concer...MECOM is a master repressor of myeloid differentiation through dose control of CEBPA in acute myeloid leukemia
Dorien Pastoors et al.
Enhancer translocations, due to 3q26 rearrangements, drive out-of-context MECOM expression in an aggressive subtype of acute myeloid leukemia (AML). Direct depletion of MECOM using an endogenous auxin-inducible degron immediately upregulates expression of myeloid differentiation factor CEBPA. MECOM depletion is also...Nuclear localization of MTHFD2 is required for correct mitosis progression
Natalia Pardo-Lorente et al.
Subcellular compartmentalization of metabolic enzymes establishes a unique metabolic environment that elicits specific cellular functions. Indeed, the nuclear translocation of certain metabolic enzymes is required for epigenetic regulation and gene expression control. Here, we show that the nuclear localization of t...Claudin-1 as a potential marker of stress-induced premature senescence in vascular smooth muscle cells
Agnieszka Gadecka et al.
Cellular senescence, a permanent state of cell cycle arrest, can result either from external stress and is then called stress-induced premature senescence (SIPS), or from the exhaustion of cell division potential giving rise to replicative senescence (RS). Despite numerous biomarkers distinguishing SIPS from RS rema...RNA sequestration in P-bodies sustains myeloid leukaemia
Srikanth Kodali et al.
Post-transcriptional mechanisms are fundamental safeguards of progenitor cell identity and are often dysregulated in cancer. Here, we identified regulators of P-bodies as crucial vulnerabilities in acute myeloid leukaemia (AML) through genome-wide CRISPR screens in normal and malignant haematopoietic progenitors. We...A multiomic atlas of the aging hippocampus reveals molecular changes in response to environmental enrichment
Perez R. F. at al.
Aging involves the deterioration of organismal function, leading to the emergence of multiple pathologies. Environmental stimuli, including lifestyle, can influence the trajectory of this process and may be used as tools in the pursuit of healthy aging. To evaluate the role of epigenetic mechanisms in this context, ...In vitro production of cat-restricted Toxoplasma pre-sexual stages
Antunes, A.V. et al.
Sexual reproduction of Toxoplasma gondii, confined to the felid gut, remains largely uncharted owing to ethical concerns regarding the use of cats as model organisms. Chromatin modifiers dictate the developmental fate of the parasite during its multistage life cycle, but their targeting to stage-specific cistro...Alterations in the hepatocyte epigenetic landscape in steatosis.
Maji Ranjan K. et al.
Fatty liver disease or the accumulation of fat in the liver, has been reported to affect the global population. This comes with an increased risk for the development of fibrosis, cirrhosis, and hepatocellular carcinoma. Yet, little is known about the effects of a diet containing high fat and alcohol towards epigenet...Chromatin profiling identifies transcriptional readthrough as a conservedmechanism for piRNA biogenesis in mosquitoes.
Qu J. et al.
The piRNA pathway in mosquitoes differs substantially from other model organisms, with an expanded PIWI gene family and functions in antiviral defense. Here, we define core piRNA clusters as genomic loci that show ubiquitous piRNA expression in both somatic and germline tissues. These core piRNA clusters are enriche...Epigenetic dosage identifies two major and functionally distinct beta cells ubtypes.
Dror E.et al.
The mechanisms that specify and stabilize cell subtypes remain poorly understood. Here, we identify two major subtypes of pancreatic β cells based on histone mark heterogeneity (beta HI and beta LO). Beta HI cells exhibit 4-fold higher levels of H3K27me3, distinct chromatin organization and compaction, a...Species-specific regulation of XIST by the JPX/FTX orthologs.
Rosspopoff O. et al.
X chromosome inactivation (XCI) is an essential process, yet it initiates with remarkable diversity in various mammalian species. XIST, the main trigger of XCI, is controlled in the mouse by an interplay of lncRNA genes (LRGs), some of which evolved concomitantly to XIST and have orthologues across all placental mam...Noncanonical regulation of imprinted gene Igf2 by amyloid-beta 1-42 inAlzheimer's disease.
Fertan E. et al.
Reduced insulin-like growth factor 2 (IGF2) levels in Alzheimer's disease (AD) may be the mechanism relating age-related metabolic disorders to dementia. Since Igf2 is an imprinted gene, we examined age and sex differences in the relationship between amyloid-beta 1-42 (Aβ) accumulation and epigenetic regulation...Histone remodeling reflects conserved mechanisms of bovine and humanpreimplantation development.
Zhou C. et al.
How histone modifications regulate changes in gene expression during preimplantation development in any species remains poorly understood. Using CUT\&Tag to overcome limiting amounts of biological material, we profiled two activating (H3K4me3 and H3K27ac) and two repressive (H3K9me3 and H3K27me3) marks in bovine...Dietary methionine starvation impairs acute myeloid leukemia progression.
Cunningham A. et al.
Targeting altered tumor cell metabolism might provide an attractive opportunity for patients with acute myeloid leukemia (AML). An amino acid dropout screen on primary leukemic stem cells and progenitor populations revealed a number of amino acid dependencies, of which methionine was one of the strongest. By using v...Dominant role of DNA methylation over H3K9me3 for IAP silencingin endoderm.
Wang Z. et al.
Silencing of endogenous retroviruses (ERVs) is largely mediated by repressive chromatin modifications H3K9me3 and DNA methylation. On ERVs, these modifications are mainly deposited by the histone methyltransferase Setdb1 and by the maintenance DNA methyltransferase Dnmt1. Knock-out of either Setdb1 or Dnmt1 leads to...bESCs from cloned embryos do not retain transcriptomic or epigenetic memory from somatic donor cells.
Navarro M. et al.
Embryonic stem cells (ESC) indefinitely maintain the pluripotent state of the blastocyst epiblast. Stem cells are invaluable for studying development and lineage commitment, and in livestock they constitute a useful tool for genomic improvement and in vitro breeding programs. Although these cells have been recently ...Epigenetic Mechanisms Mediating Cell State Transitions in Chondrocytes
Wuelling M. et al.
Epigenetic modifications play critical roles in regulating cell lineage differentiation, but the epigenetic mechanisms guiding specific differentiation steps within a cell lineage have rarely been investigated. To decipher such mechanisms, we used the defined transition from proliferating (PC) into hypertrophic chon...Comprehensive characterization of the epigenetic landscape in Multiple Myeloma
Elina Alaterre et al.
Background: Human multiple myeloma (MM) cell lines (HMCLs) have been widely used to understand themolecular processes that drive MM biology. Epigenetic modifications are involved in MM development,progression, and drug resistance. A comprehensive characterization of the epigenetic landscape of MM wouldadvance our un...Comprehensive characterization of the epigenetic landscape in Multiple Myeloma
Alaterre, Elina and Ovejero, Sara and Herviou, Laurie and de Boussac, Hugues and Papadopoulos, Giorgio and Kulis, Marta and Boireau, Stéphanie and Robert, Nicolas and Requirand, Guilhem and Bruyer, Angélique and Cartron, Guillaume and Vincent, Laure and M
Background: Human multiple myeloma (MM) cell lines (HMCLs) have been widely used to understand the molecular processes that drive MM biology. Epigenetic modifications are involved in MM development, progression, and drug resistance. A comprehensive characterization of the epigenetic landscape of MM would advance our...Enhanced targeted DNA methylation of the CMV and endogenous promoterswith dCas9-DNMT3A3L entails distinct subsequent histonemodification changes in CHO cells.
Marx Nicolas et al.
With the emergence of new CRISPR/dCas9 tools that enable site specific modulation of DNA methylation and histone modifications, more detailed investigations of the contribution of epigenetic regulation to the precise phenotype of cells in culture, including recombinant production subclones, is now possible. These al...Lasp1 regulates adherens junction dynamics and fibroblast transformationin destructive arthritis
Beckmann D. et al.
The LIM and SH3 domain protein 1 (Lasp1) was originally cloned from metastatic breast cancer and characterised as an adaptor molecule associated with tumourigenesis and cancer cell invasion. However, the regulation of Lasp1 and its function in the aggressive transformation of cells is unclear. Here we use integrativ...Sarcomere function activates a p53-dependent DNA damage response that promotes polyploidization and limits in vivo cell engraftment.
Pettinato, Anthony M. et al.
Human cardiac regeneration is limited by low cardiomyocyte replicative rates and progressive polyploidization by unclear mechanisms. To study this process, we engineer a human cardiomyocyte model to track replication and polyploidization using fluorescently tagged cyclin B1 and cardiac troponin T. Using time-lapse i...Androgen and glucocorticoid receptor direct distinct transcriptionalprograms by receptor-specific and shared DNA binding sites.
Kulik, Marina et al.
The glucocorticoid (GR) and androgen (AR) receptors execute unique functions in vivo, yet have nearly identical DNA binding specificities. To identify mechanisms that facilitate functional diversification among these transcription factor paralogs, we studied them in an equivalent cellular context. Analysis of chroma...Environmental enrichment induces epigenomic and genome organization changesrelevant for cognitive function
Espeso-Gil, S. et al.
In early development, the environment triggers mnemonic epigenomic programs resulting in memory and learning experiences to confer cognitive phenotypes into adulthood. To uncover how environmental stimulation impacts the epigenome and genome organization, we used the paradigm of environmental enrichment (EE) in youn...TRF2 Mediates Replication Initiation within Human Telomeres to PreventTelomere Dysfunction.
Drosopoulos, William C and Deng, Zhong and Twayana, Shyam and Kosiyatrakul,Settapong T and Vladimirova, Olga and Lieberman, Paul M and Schildkraut,Carl L
The telomeric shelterin protein telomeric repeat-binding factor 2 (TRF2) recruits origin recognition complex (ORC) proteins, the foundational building blocks of DNA replication origins, to telomeres. We seek to determine whether TRF2-recruited ORC proteins give rise to functional origins in telomere repeat tracts. W...β-Glucan Induces Protective Trained Immunity against Mycobacterium tuberculosis Infection: A Key Role for IL-1.
Moorlag SJCFM, Khan N, Novakovic B, Kaufmann E, Jansen T, van Crevel R, Divangahi M, Netea MG
β-glucan is a potent inducer of epigenetic and functional reprogramming of innate immune cells, a process called "trained immunity," resulting in an enhanced host response against secondary infections. We investigate whether β-glucan exposure confers protection against pulmonary Mycobacterium tuberculosis ...Endogenous retroviruses are a source of enhancers with oncogenic potential in acute myeloid leukaemia
/
Acute myeloid leukemia (AML) is a highly aggressive hematopoietic malignancy, defined by a series of genetic and epigenetic alterations, which result in deregulation of transcriptional networks. One understudied but important source of transcriptional regulators are transposable elements (TEs), which are widespread ...Targeting Macrophage Histone H3 Modification as a Leishmania Strategy to Dampen the NF-κB/NLRP3-Mediated Inflammatory Response.
Lecoeur H, Prina E, Rosazza T, Kokou K, N'Diaye P, Aulner N, Varet H, Bussotti G, Xing Y, Milon G, Weil R, Meng G, Späth GF
Aberrant macrophage activation during intracellular infection generates immunopathologies that can cause severe human morbidity. A better understanding of immune subversion strategies and macrophage phenotypic and functional responses is necessary to design host-directed intervention strategies. Here, we uncover a f...Analysis of Histone Modifications in Rodent Pancreatic Islets by Native Chromatin Immunoprecipitation.
Sandovici I, Nicholas LM, O'Neill LP
The islets of Langerhans are clusters of cells dispersed throughout the pancreas that produce several hormones essential for controlling a variety of metabolic processes, including glucose homeostasis and lipid metabolism. Studying the transcriptional control of pancreatic islet cells has important implications for ...Mkt1 is required for RNAi-mediated silencing and establishment of heterochromatin in fission yeast.
Taglini F, Chapman E, van Nues R, Theron E, Bayne EH
Constitutive domains of repressive heterochromatin are maintained within the fission yeast genome through self-reinforcing mechanisms involving histone methylation and small RNAs. Non-coding RNAs generated from heterochromatic regions are processed into small RNAs by the RNA interference pathway, and are subject to ...β-Glucan-Induced Trained Immunity Protects against Leishmania braziliensis Infection: a Crucial Role for IL-32.
Dos Santos JC, Barroso de Figueiredo AM, Teodoro Silva MV, Cirovic B, de Bree LCJ, Damen MSMA, Moorlag SJCFM, Gomes RS, Helsen MM, Oosting M, Keating ST, Schlitzer A, Netea MG, Ribeiro-Dias F, Joosten LAB
American tegumentary leishmaniasis is a vector-borne parasitic disease caused by Leishmania protozoans. Innate immune cells undergo long-term functional reprogramming in response to infection or Bacillus Calmette-Guérin (BCG) vaccination via a process called trained immunity, conferring non-specific protectio...Development and epigenetic plasticity of murine Müller glia.
Dvoriantchikova G, Seemungal RJ, Ivanov D
The ability to regenerate the entire retina and restore lost sight after injury is found in some species and relies mostly on the epigenetic plasticity of Müller glia. To understand the role of mammalian Müller glia as a source of progenitors for retinal regeneration, we investigated changes in gene expres...Probing the Tumor Suppressor Function of BAP1 in CRISPR-Engineered Human Liver Organoids.
Artegiani B, van Voorthuijsen L, Lindeboom RGH, Seinstra D, Heo I, Tapia P, López-Iglesias C, Postrach D, Dayton T, Oka R, Hu H, van Boxtel R, van Es JH, Offerhaus J, Peters PJ, van Rheenen J, Vermeulen M, Clevers H
The deubiquitinating enzyme BAP1 is a tumor suppressor, among others involved in cholangiocarcinoma. BAP1 has many proposed molecular targets, while its Drosophila homolog is known to deubiquitinate histone H2AK119. We introduce BAP1 loss-of-function by CRISPR/Cas9 in normal human cholangiocyte organoids. We find th...The epigenetic basis for the impaired ability of adult murine retinal pigment epithelium cells to regenerate retinal tissue.
Dvoriantchikova G, Seemungal RJ, Ivanov D
The epigenetic plasticity of amphibian retinal pigment epithelium (RPE) allows them to regenerate the entire retina, a trait known to be absent in mammals. In this study, we investigated the epigenetic plasticity of adult murine RPE to identify possible mechanisms that prevent mammalian RPE from regenerating retinal...CBX7 Induces Self-Renewal of Human Normal and Malignant Hematopoietic Stem and Progenitor Cells by Canonical and Non-canonical Interactions.
Jung J, Buisman SC, Weersing E, Dethmers-Ausema A, Zwart E, Schepers H, Dekker MR, Lazare SS, Hammerl F, Skokova Y, Kooistra SM, Klauke K, Poot RA, Bystrykh LV, de Haan G
In this study, we demonstrate that, among all five CBX Polycomb proteins, only CBX7 possesses the ability to control self-renewal of human hematopoietic stem and progenitor cells (HSPCs). Xenotransplantation of CBX7-overexpressing HSPCs resulted in increased multi-lineage long-term engraftment and myelopoiesis. ...Chromatin-Based Classification of Genetically Heterogeneous AMLs into Two Distinct Subtypes with Diverse Stemness Phenotypes.
Yi G, Wierenga ATJ, Petraglia F, Narang P, Janssen-Megens EM, Mandoli A, Merkel A, Berentsen K, Kim B, Matarese F, Singh AA, Habibi E, Prange KHM, Mulder AB, Jansen JH, Clarke L, Heath S, van der Reijden BA, Flicek P, Yaspo ML, Gut I, Bock C, Schuringa JJ
Global investigation of histone marks in acute myeloid leukemia (AML) remains limited. Analyses of 38 AML samples through integrated transcriptional and chromatin mark analysis exposes 2 major subtypes. One subtype is dominated by patients with NPM1 mutations or MLL-fusion genes, shows activation of the regulat...Angiotensin II induced CSF1 transcription is mediated by a crosstalk between different epigenetic factors in vascular endothelial cells.
Shao J, Weng X, Zhuo L, Yu L, Li Z, Shen K, Xu W, Fang M, Xu Y
Endothelium-derived colony stimulating factor (CSF1) plays a key role in a range of human pathologies. Angiotensin II (Ang II) has been documented to stimulate CSF1 transcription although the underlying epigenetic mechanism remains unclear. Here we report that induction of CSF1 transcription by Ang II in vascular en...The Itaconate Pathway Is a Central Regulatory Node Linking Innate Immune Tolerance and Trained Immunity
Domínguez-Andrés Jorge, Novakovic Boris, Li Yang, Scicluna Brendon P., Gresnigt Mark S., Arts Rob J.W., Oosting Marije, Moorlag Simone J.C.F.M., Groh Laszlo A., Zwaag Jelle, Koch Rebecca M., ter Horst Rob, Joosten Leo A.B., Wijmenga Cisca, Michelucci Ales
Sepsis involves simultaneous hyperactivation of the immune system and immune paralysis, leading to both organ dysfunction and increased susceptibility to secondary infections. Acute activation of myeloid cells induced itaconate synthesis, which subsequently mediated innate immune tolerance in human monocytes. In con...The Polycomb-Dependent Epigenome Controls β Cell Dysfunction, Dedifferentiation, and Diabetes.
Lu TT, Heyne S, Dror E, Casas E, Leonhardt L, Boenke T, Yang CH, Sagar , Arrigoni L, Dalgaard K, Teperino R, Enders L, Selvaraj M, Ruf M, Raja SJ, Xie H, Boenisch U, Orkin SH, Lynn FC, Hoffman BG, Grün D, Vavouri T, Lempradl AM, Pospisilik JA
To date, it remains largely unclear to what extent chromatin machinery contributes to the susceptibility and progression of complex diseases. Here, we combine deep epigenome mapping with single-cell transcriptomics to mine for evidence of chromatin dysregulation in type 2 diabetes. We find two chromatin-state signat...The reference epigenome and regulatory chromatin landscape of chronic lymphocytic leukemia
Beekman R. et al.
Chronic lymphocytic leukemia (CLL) is a frequent hematological neoplasm in which underlying epigenetic alterations are only partially understood. Here, we analyze the reference epigenome of seven primary CLLs and the regulatory chromatin landscape of 107 primary cases in the context of normal B cell differentiation....Increased H3K9 methylation and impaired expression of Protocadherins are associated with the cognitive dysfunctions of the Kleefstra syndrome.
Iacono G, Dubos A, Méziane H, Benevento M, Habibi E, Mandoli A, Riet F, Selloum M, Feil R, Zhou H, Kleefstra T, Kasri NN, van Bokhoven H, Herault Y, Stunnenberg HG
Kleefstra syndrome, a disease with intellectual disability, autism spectrum disorders and other developmental defects is caused in humans by haploinsufficiency of EHMT1. Although EHMT1 and its paralog EHMT2 were shown to be histone methyltransferases responsible for deposition of the di-methylated H3K9 (H3K9me2), th...The Ftx Noncoding Locus Controls X Chromosome Inactivation Independently of Its RNA Products.
Furlan G, Gutierrez Hernandez N, Huret C, Galupa R, van Bemmel JG, Romito A, Heard E, Morey C, Rougeulle C
Accumulation of the Xist long noncoding RNA (lncRNA) on one X chromosome is the trigger for X chromosome inactivation (XCI) in female mammals. Xist expression, which needs to be tightly controlled, involves a cis-acting region, the X-inactivation center (Xic), containing many lncRNA genes that evolved concomita...Metabolic Induction of Trained Immunity through the Mevalonate Pathway.
Bekkering S, Arts RJW, Novakovic B, Kourtzelis I, van der Heijden CDCC, Li Y, Popa CD, Ter Horst R, van Tuijl J, Netea-Maier RT, van de Veerdonk FL, Chavakis T, Joosten LAB, van der Meer JWM, Stunnenberg H, Riksen NP, Netea MG
Innate immune cells can develop long-term memory after stimulation by microbial products during infections or vaccinations. Here, we report that metabolic signals can induce trained immunity. Pharmacological and genetic experiments reveal that activation of the cholesterol synthesis pathway, but not the synthesis of...BCG Vaccination Protects against Experimental Viral Infection in Humans through the Induction of Cytokines Associated with Trained Immunity.
Arts RJW, Moorlag SJCFM, Novakovic B, Li Y, Wang SY, Oosting M, Kumar V, Xavier RJ, Wijmenga C, Joosten LAB, Reusken CBEM, Benn CS, Aaby P, Koopmans MP, Stunnenberg HG, van Crevel R, Netea MG
The tuberculosis vaccine bacillus Calmette-Guérin (BCG) has heterologous beneficial effects against non-related infections. The basis of these effects has been poorly explored in humans. In a randomized placebo-controlled human challenge study, we found that BCG vaccination induced genome-wide epigenetic repr...DNA methylation signatures follow preformed chromatin compartments in cardiac myocytes
Nothjunge S. et al.
Storage of chromatin in restricted nuclear space requires dense packing while ensuring DNA accessibility. Thus, different layers of chromatin organization and epigenetic control mechanisms exist. Genome-wide chromatin interaction maps revealed large interaction domains (TADs) and higher order A and B compartments, r...Chromosome contacts in activated T cells identify autoimmune disease candidate genes
Burren OS et al.
BACKGROUND: Autoimmune disease-associated variants are preferentially found in regulatory regions in immune cells, particularly CD4+ T cells. Linking such regulatory regions to gene promoters in disease-relevant cell contexts facilitates identification of candidate disease genes. RESULTS: Within 4 h, act...DNA methylation heterogeneity defines a disease spectrum in Ewing sarcoma
Sheffield N.C. et al.
Developmental tumors in children and young adults carry few genetic alterations, yet they have diverse clinical presentation. Focusing on Ewing sarcoma, we sought to establish the prevalence and characteristics of epigenetic heterogeneity in genetically homogeneous cancers. We performed genome-scale DNA methylation ...Immunometabolic Pathways in BCG-Induced Trained Immunity
Arts R.J. et al.
The protective effects of the tuberculosis vaccine Bacillus Calmette-Guerin (BCG) on unrelated infections are thought to be mediated by long-term metabolic changes and chromatin remodeling through histone modifications in innate immune cells such as monocytes, a process termed trained immunity. Here, we show that BC...TET-dependent regulation of retrotransposable elements in mouse embryonic stem cells
de la Rica L. et al.
BACKGROUND: Ten-eleven translocation (TET) enzymes oxidise DNA methylation as part of an active demethylation pathway. Despite extensive research into the role of TETs in genome regulation, little is known about their effect on transposable elements (TEs), which make up nearly half of the mouse and human genomes....β-Glucan Reverses the Epigenetic State of LPS-Induced Immunological Tolerance
Novakovic B. et al.
Innate immune memory is the phenomenon whereby innate immune cells such as monocytes or macrophages undergo functional reprogramming after exposure to microbial components such as lipopolysaccharide (LPS). We apply an integrated epigenomic approach to characterize the molecular events involved in LPS-induced to...The Hematopoietic Transcription Factors RUNX1 and ERG Prevent AML1-ETO Oncogene Overexpression and Onset of the Apoptosis Program in t(8;21) AMLs
Mandoli A. et al.
The t(8;21) acute myeloid leukemia (AML)-associated oncoprotein AML1-ETO disrupts normal hematopoietic differentiation. Here, we have investigated its effects on the transcriptome and epigenome in t(8,21) patient cells. AML1-ETO binding was found at promoter regions of active genes with high levels of histone acetyl...Neonatal monocytes exhibit a unique histone modification landscape
Bermick JR et al.
Background Neonates have dampened expression of pro-inflammatory cytokines and difficulty clearing pathogens. This makes them uniquely susceptible to infections, but the factors regulating neonatal-specific immune responses are poorly understood. Epigenetics, including histone modifications, can activate or silen...Epigenetic dynamics of monocyte-to-macrophage differentiation
Wallner S et al.
BACKGROUND: Monocyte-to-macrophage differentiation involves major biochemical and structural changes. In order to elucidate the role of gene regulatory changes during this process, we used high-throughput sequencing to analyze the complete transcriptome and epigenome of human monocytes that were differentiated in...Comprehensive genome and epigenome characterization of CHO cells in response to evolutionary pressures and over time
Feichtinger J, Hernández I, Fischer C, Hanscho M, Auer N, Hackl M, Jadhav V, Baumann M, Krempl PM, Schmidl C, Farlik M, Schuster M, Merkel A, Sommer A, Heath S, Rico D, Bock C, Thallinger GG, Borth N
The most striking characteristic of CHO cells is their adaptability, which enables efficient production of proteins as well as growth under a variety of culture conditions, but also results in genomic and phenotypic instability. To investigate the relative contribution of genomic and epigenetic modifications towards...Deciphering the principles that govern mutually exclusive expression of Plasmodium falciparum clag3 genes
Rovira-Graells N, Crowley VM, Bancells C, Mira-Martínez S, de Pouplana LR, Cortés A
The product of the Plasmodium falciparum genes clag3.1 and clag3.2 plays a fundamental role in malaria parasite biology by determining solute transport into infected erythrocytes. Expression of the two clag3 genes is mutually exclusive, such that a single parasite expresses only one of the two genes at a time. Here ...Epigenome mapping reveals distinct modes of gene regulation and widespread enhancer reprogramming by the oncogenic fusion protein EWS-FLI1.
Tomazou EM, Sheffield NC, Schmidl C, Schuster M, Schönegger A, Datlinger P, Kubicek S, Bock C, Kovar H
Transcription factor fusion proteins can transform cells by inducing global changes of the transcriptome, often creating a state of oncogene addiction. Here, we investigate the role of epigenetic mechanisms in this process, focusing on Ewing sarcoma cells that are dependent on the EWS-FLI1 fusion protein. We establi... - 関連商品


