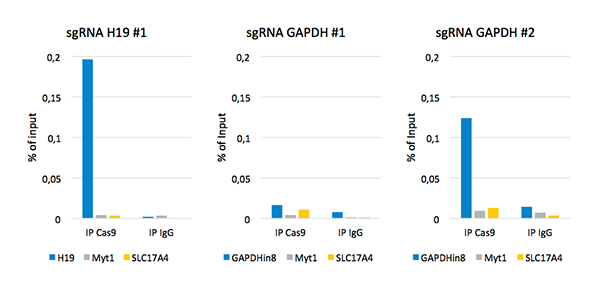
ChIP was performed on sheared chromatin from 4,000,000 HEK293T cells using the iDeal ChIP-seq Kit for Transcription Factors, 5 µl of the polyclonal Cas9 antibody and 1 µg of the negative control IgG. Primers specific for the human H19, GAPDH, Myt1 and SLC17A4 were used for the qPCR. The figure shows the recovery, expressed as a percent of input (the relative amount of immunoprecipitated DNA compared to input after qPCR analysis).
Genome editing (CRISPR/Cas9)
Selection of guide RNA by ChIP
While a number of computational tools help design sgRNAs to target specific loci, the accurate prediction of whether Cas9 and the sgRNA of interest will bind specifically to the genome is still a challenge.
In order to ensure specific targeting of CRISPR/Cas9, the verification of the specific binding of the sgRNA at the locus of interest is required. The methodology of choice for such verification is chromatin immunoprecipitation followed either by real-time PCR (ChIP-qPCR) or sequencing (ChIP-seq) to analyze protein-DNA interactions.
Diagenode’s iDeal ChIP-seq Kit for Transcription Factors provides a robust workflow to investigate dCas9 binding in the genome by specifically enriching dCas9 in the on-target region by ChIP followed by real-time PCR. In addition, sequencing analysis of the DNA immunoprecipitated with the Cas9 antibody illustrates potential genome-wide off-target effects. This information is essential to verify the correct binding of dCas9 fused to an effector.
1. Monitoring of sgRNA efficiency by ChIP-qPCR
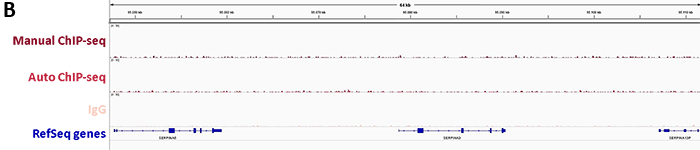
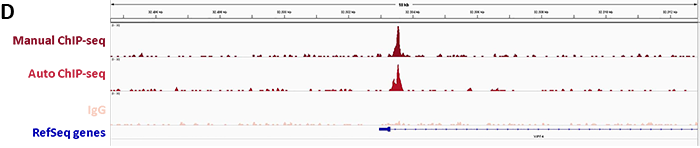
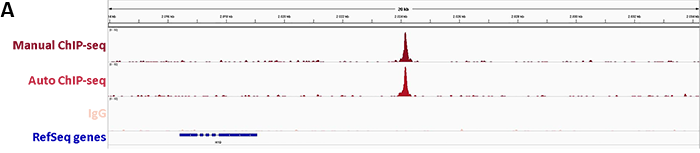
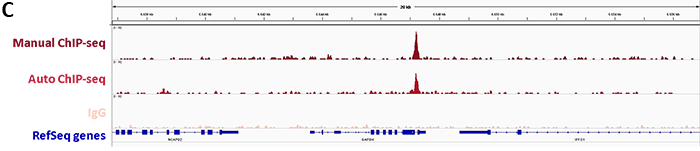
2. Confirmation of binding specificity of sgRNA and detection of off-target bindings by ChIP-seq