| ELISA Enzyme-linked immunosorbent assay. Read more |
| DB Dot blotting Read more |
| WB Western blot : The quality of antibodies used in this technique is crucial for correct and specific protein identification. Diagenode offers huge selection of highly sensitive and specific western blot-validated antibodies. Learn more about: Load... Read more |
| IF Immunofluorescence: Diagenode offers huge selection of highly sensitive antibodies validated in IF. Immunofluorescence using the Diagenode monoclonal antibody directed against CRISPR/Cas9 HeLa cells transfected with a Cas9 expression vector (... Read more |
| ChIP-seq (ab) Read more |
| ChIP-qPCR (ab) Read more |
| CUT&Tag Read more |
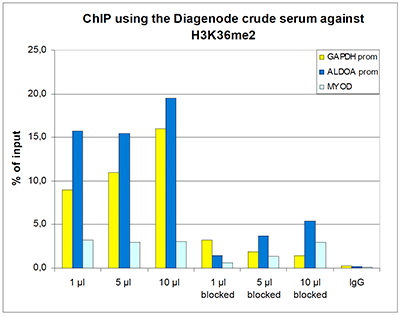
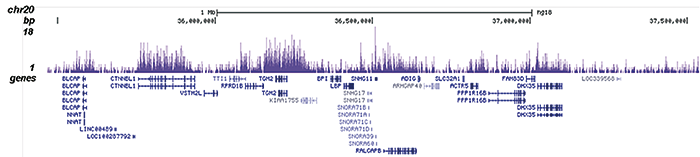
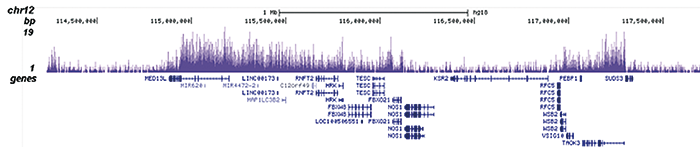
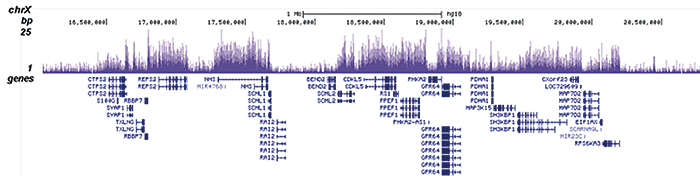
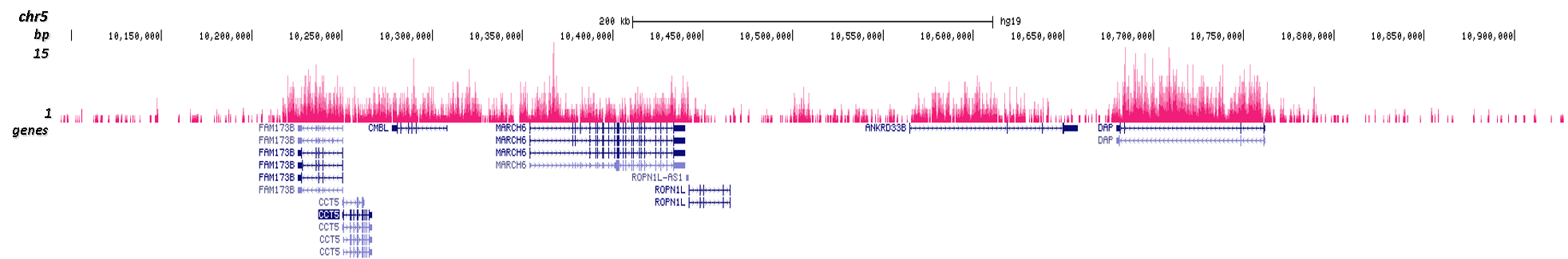
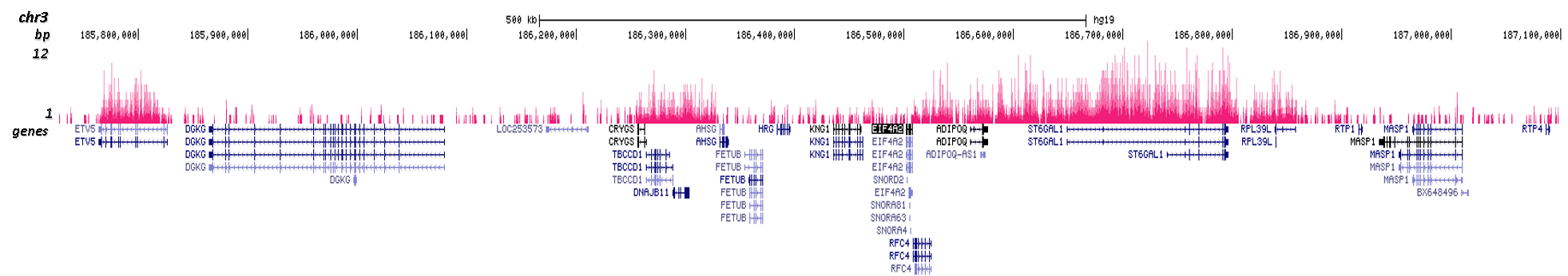
H3K36me2 Antibody - ChIP-seq Grade
| Lot | A239-001 |
|---|---|
| Concentration | Not determined |
| Species reactivity | Human, mouse, yeast: positive. Other species: not tested. |
| Type | Polyclonal |
| Purity | Whole antiserum from rabbit. |
| Host | Rabbit |
| Storage Conditions | Store at -20°C; for long storage, store at -80°C. Avoid multiple freeze-thaw cycles. |
| Storage Buffer | Whole antiserum from rabbit containing 0.05% azide |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Applications | Suggested dilution | References |
|---|---|---|
| ChIP/ChIP-seq * | 0.5-1 µl/ChIP | Fig 1, 2 |
| CUT&TAG | 1 µg | Fig 3 |
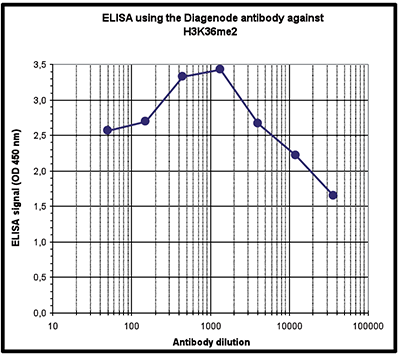
| ELISA | 1:1,000 | Fig 4 |
| Dot Blotting | 1:100,000 | Fig 5 |
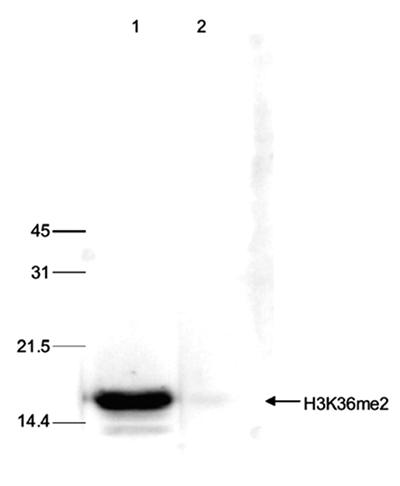
| Western Blotting | 1:1,000 | Fig 6 |
| Immunofluorescence | 1:500 | Fig 7 |
* Please note that the optimal antibody amount per IP should be determined by the end-user. We recommend testing 0.5-10 µl per IP.