How Do UMIs Work?
UMIs incorporate a unique barcode onto each molecule within a given RRBS library. By incorporating individual barcodes on each original DNA fragment, variant alleles present in the original DNA sample (true variants) can be distinguished from errors introduced during library preparation or sequencing.
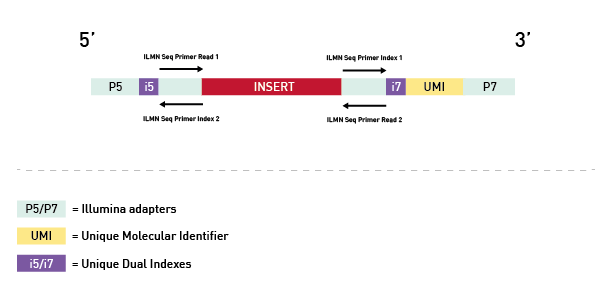
Figure 1. Premium RRBS V2 construct. Integrated UMIs enable removal of PCR duplicates during the data analysis.


