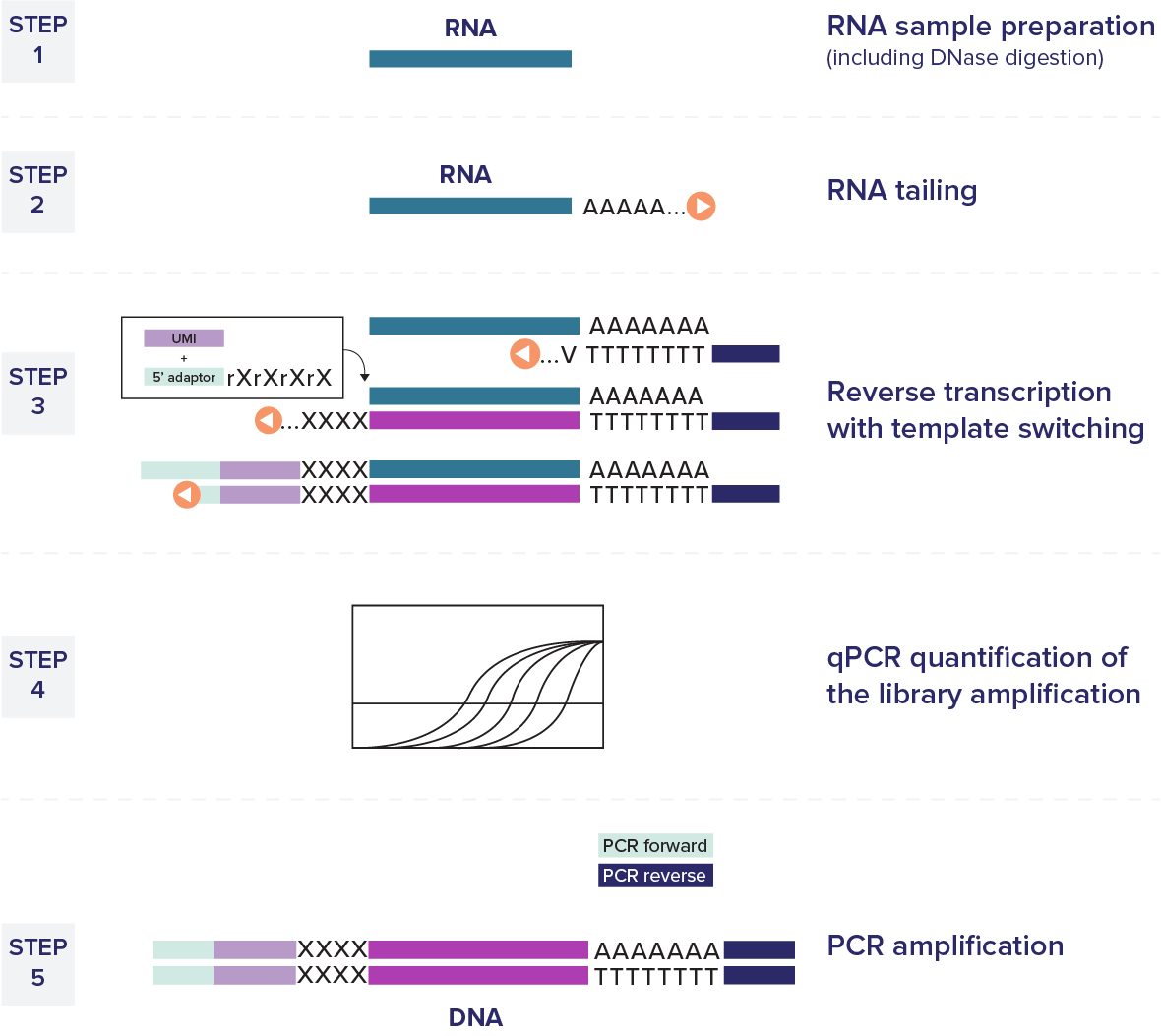
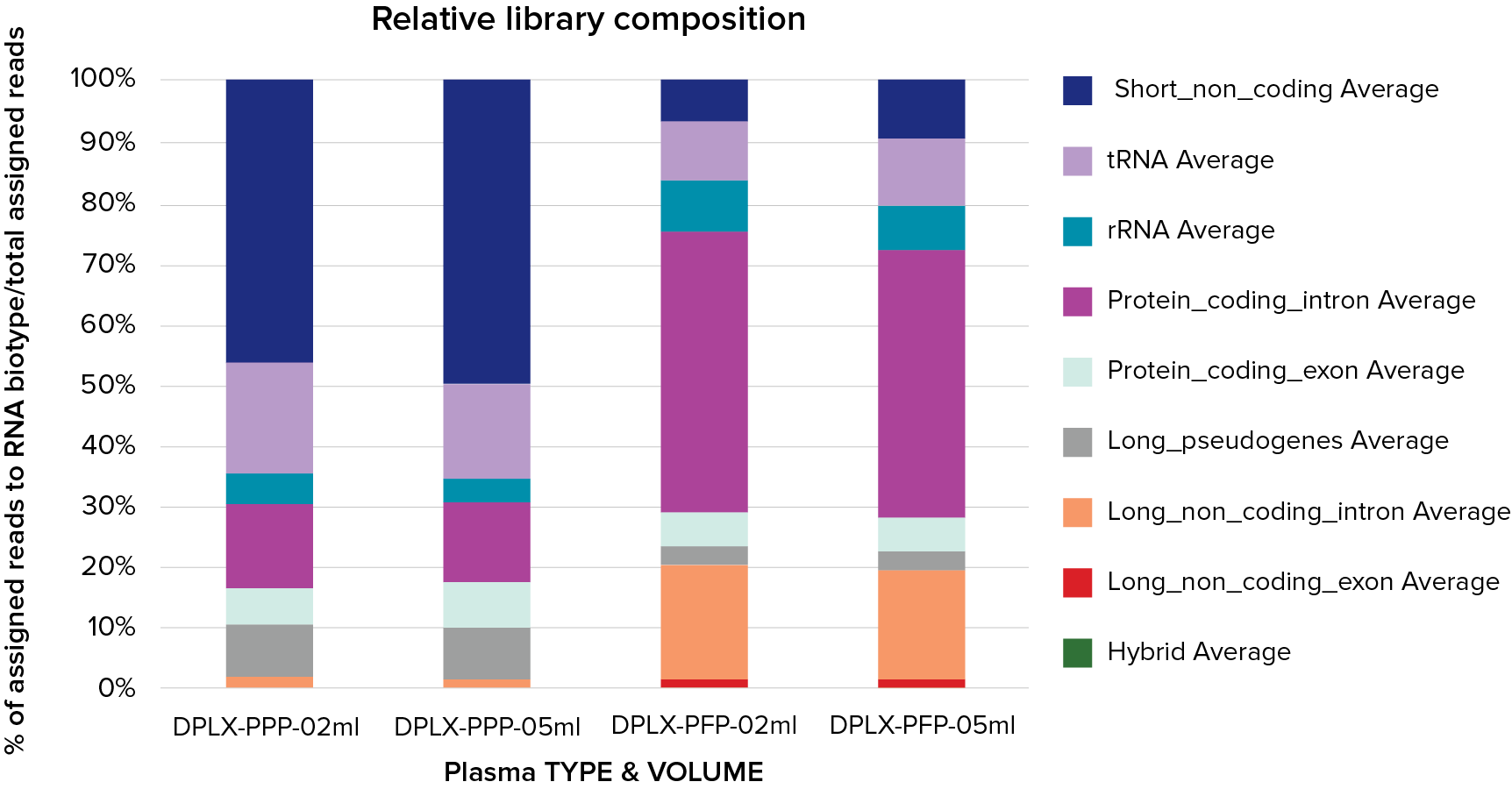The D-Plex cell-free RNA-seq technology effectively captures a broad spectrum of circulating RNAs in biofluid samples.
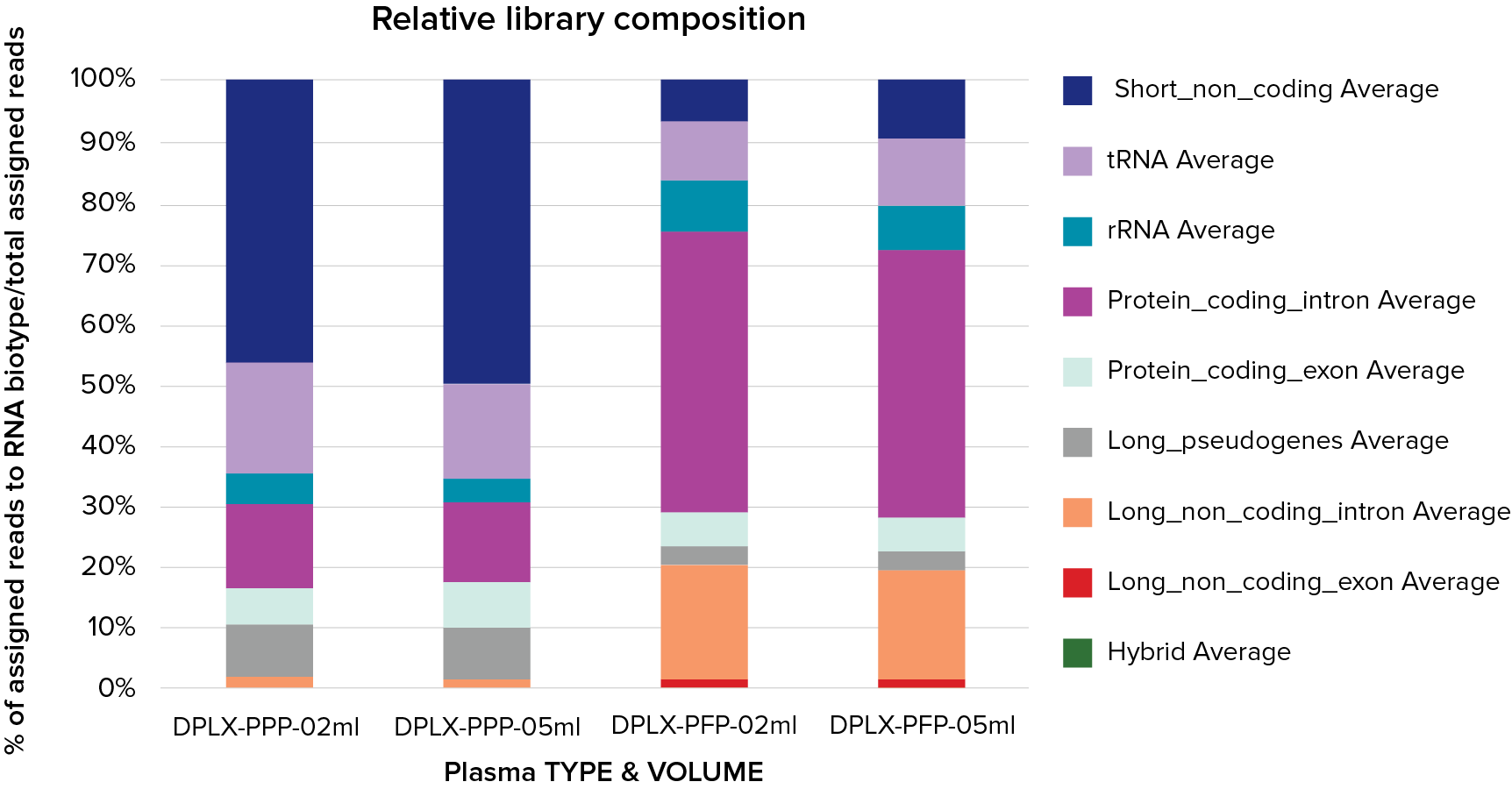
Figure 1. Typical composition of D-Plex cell-free RNA-seq libraries generated from cfRNA purified from 0.5 mL and 0.2 mL of platelet-poor plasma (PPP) and platelet-free plasma (PFP). Each condition was prepared in technical triplicates (n=3). Library composition is strongly influenced by plasma type (PPP vs PFP) and only minimally affected by the starting plasma volume, indicating that D-Plex cfRNA-seq libraries directly reflect the original cfRNA content of the sample. PPP and PFP were derived from the same pool of healthy donor blood, underscoring the critical impact of pre-analytical factors - particularly plasma preparation - on downstream data content and quality.
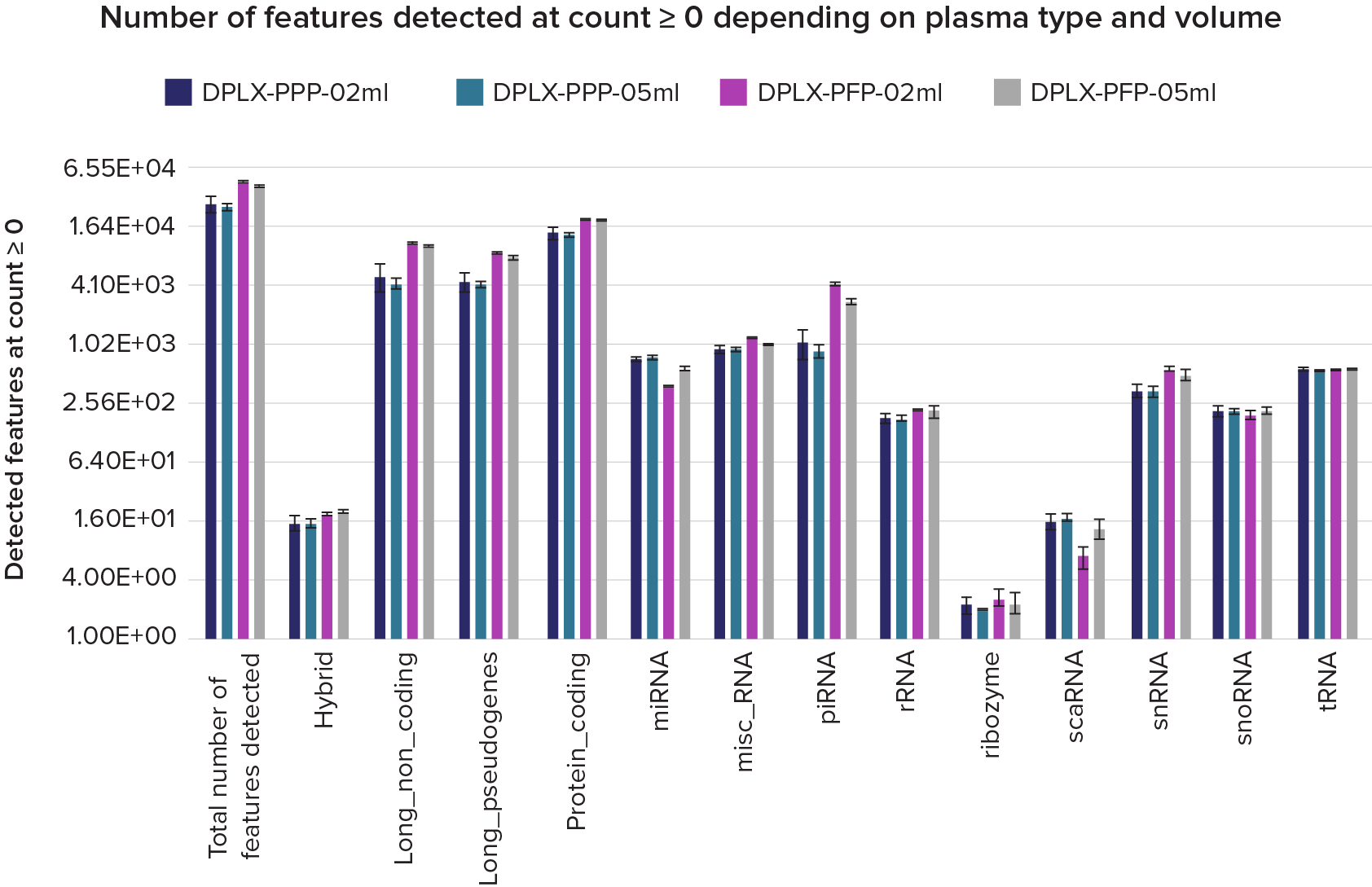
Figure 2. Detection of features with counts greater than 0 in D-Plex cell-free RNA libraries prepared from 0.5 mL and 0.2 mL of platelet-poor plasma (PPP) and platelet-free plasma (PFP). A typical D-Plex dataset provides broad coverage of the extracellular transcriptome, including protein-coding genes, long noncoding RNAs, and small noncoding RNAs such as miRNAs, snRNAs, and snoRNAs. This comprehensive feature representation makes D-Plex well suited for discovery-grade biomarker screening in translational clinical research.
D-Plex cell-free RNA-seq accurately reflects the fragmentation pattern of circulating plasma transcripts.
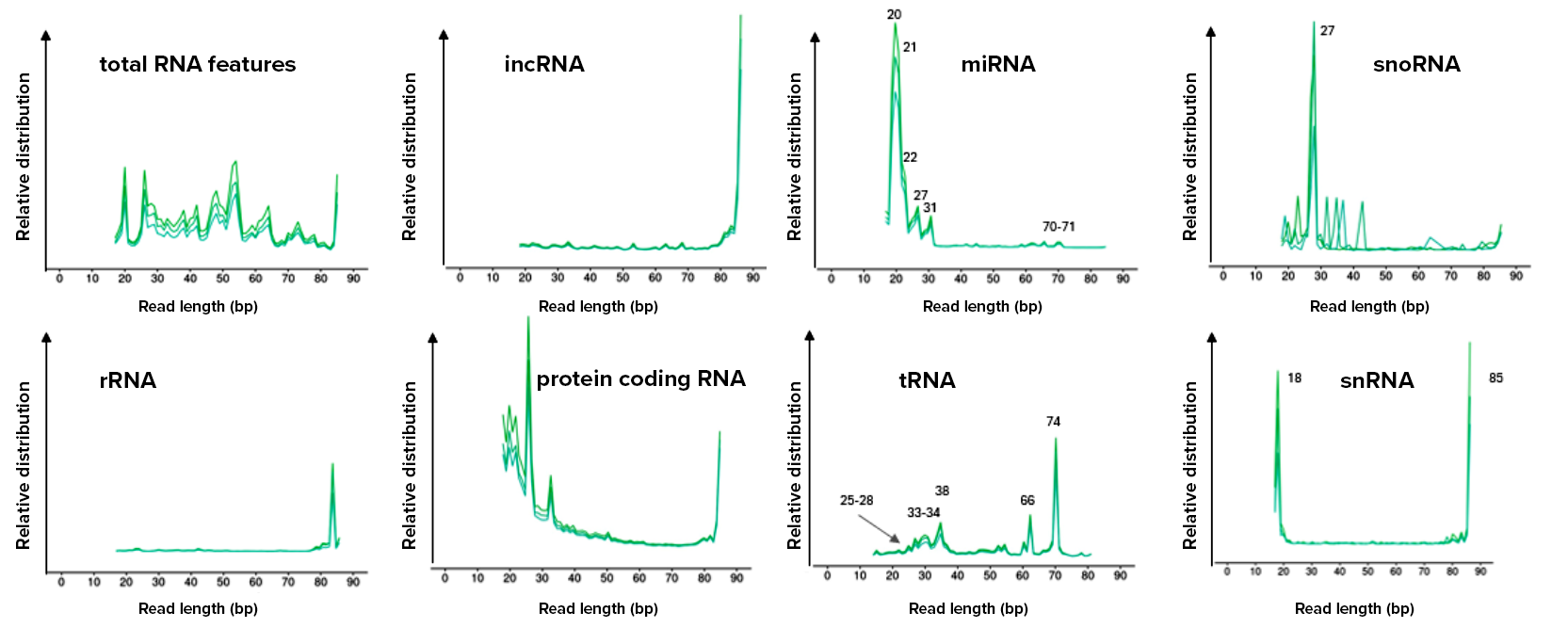
Figure 3. Fragmentation profiles of multiple RNA biotypes captured by D-Plex cell-free RNA-seq in platelet-poor plasma (PPP). D-Plex libraries recapitulate the native fragmentation patterns of cfRNA present in the original sample, enabling downstream analyses such as fragmentomics. In particular, tRNA molecules circulate predominantly as processed tRNA-derived fragments (tRFs), whose sizes align with the fragmentation profiles observed in the D-Plex data. Features longer than the maximum read length (85 bp after trimming) could not be fully measured.
D-Plex cell-free RNA-seq is a highly sensitive method that delivers consistent results across typical quantities of cell-free RNA.
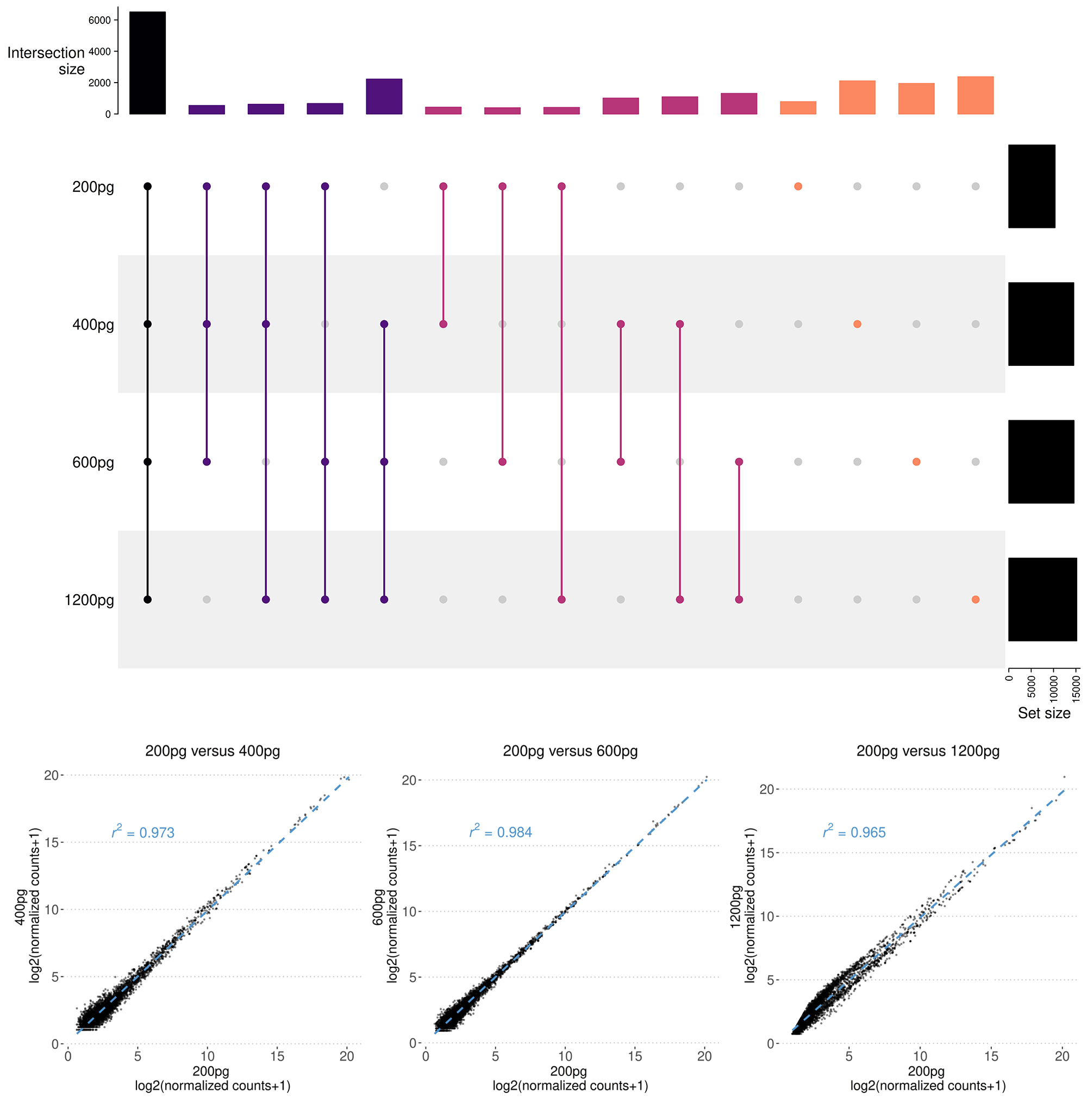
Figure 4. D-Plex cfRNA-seq consistently delivers reliable results across typical quantities of cell-free RNA used for library preparation, confirming its suitability for studying circulating RNAs in biofluids. Upset plots were created based on features identified in all samples. A feature is deemed detected if its count exceeds zero in technical triplicates for each input amount (200 pg, 400 pg, 600 pg, or 1200 pg). Scatter plots were generated using the log2 of normalized counts of the commonly detected features between the 200 pg input and each of the higher input amounts. Comparable correlation values were observed across all other input amount comparisons (not shown).
D-Plex cell-free RNA-seq demonstrates high robustness and reproducibility.
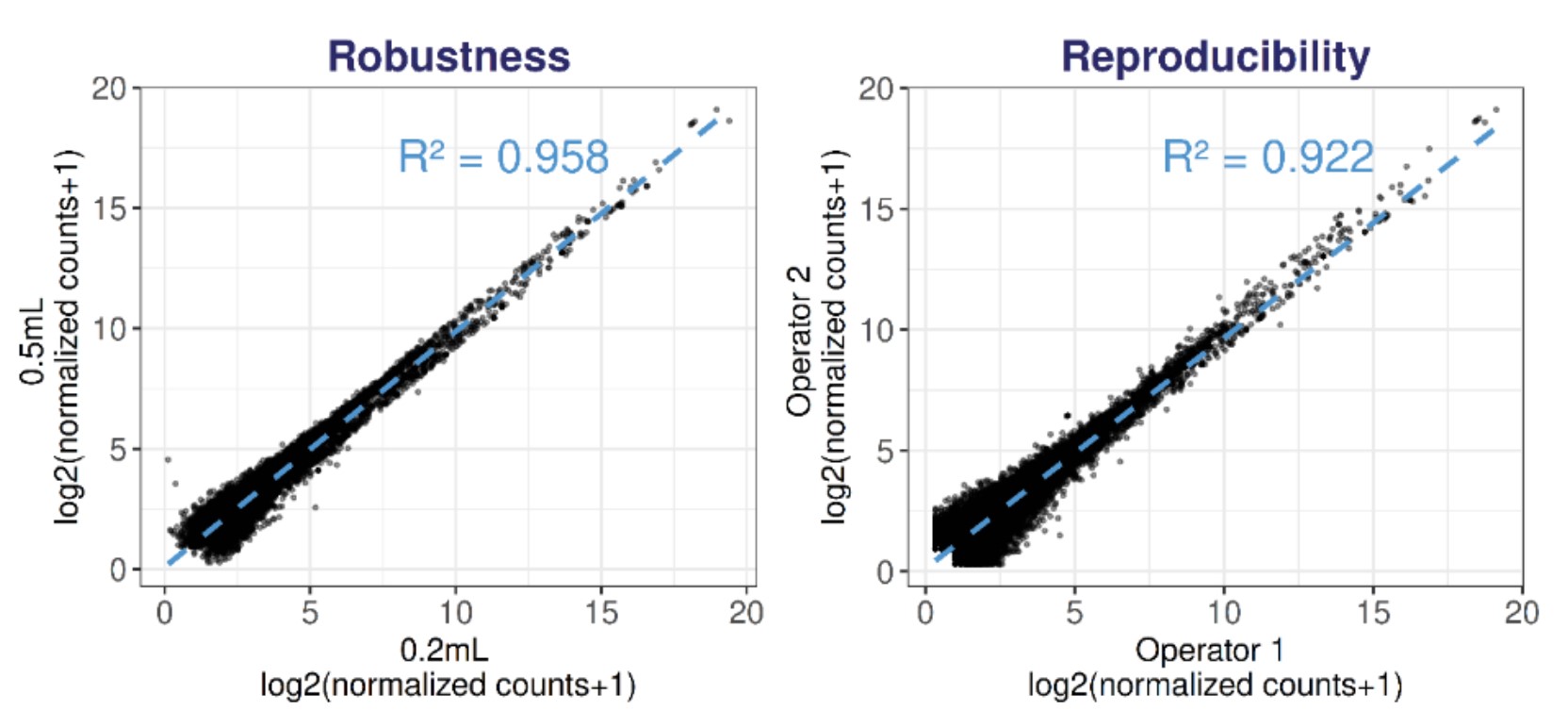
Figure 5. Robustness and reproducibility of D-Plex cell-free RNA-seq across plasma volumes and operators. (A) Robustness: Scatter plot comparing normalized counts across different plasma volumes for features commonly detected in all technical triplicates from all three operators for each plasma volume (n=9). (B) Reproducibility: Representative scatter plot comparing normalized counts across two operators at 0.5 mL input for features detected in all technical triplicates from each operator. Average pairwise correlation between operators: 0.922 at 0.5 mL and 0.866 at 0.2 mL.
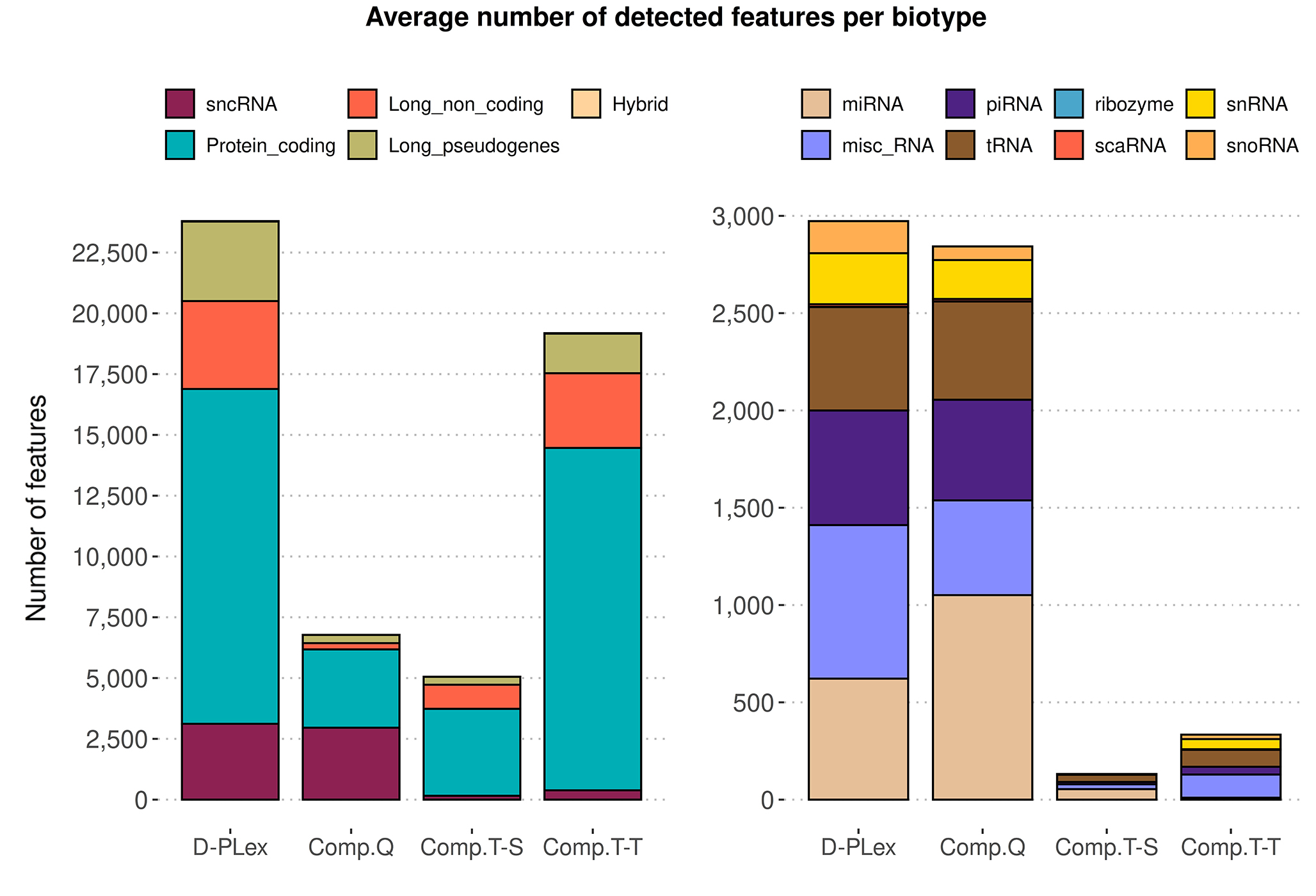
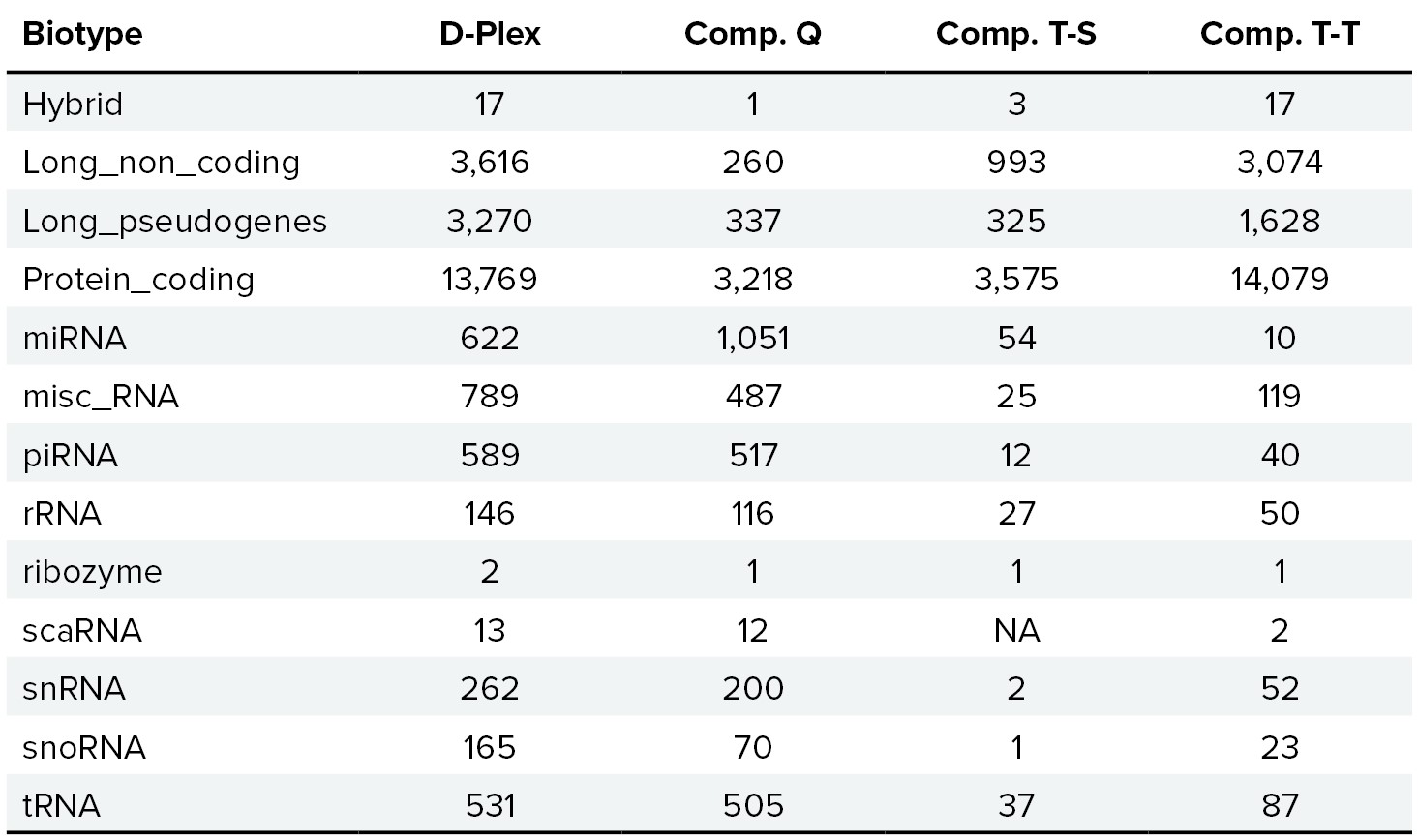
D-Plex cfRNA-seq captures a broader spectrum of cell-free RNA in plasma compared to alternative methods.
Figure 6. Unlike other library preparation kits, the D-Plex cfRNA-seq kit captures a broader spectrum of extracellular transcripts in biofluid samples, maximizing the amount of biological information available. This approach results in the highest number of detected features and the greatest biotype diversity, showcasing the powerful discovery capabilities of the D-Plex cfRNA-seq. Average number of features detected from 0.2 mL of plasma, by biotype and technology. A feature is considered detected if it shows a nonzero count across all technical triplicates from three different operators for each technology (n=9).
D-Plex cell-free RNA-seq offers superior accuracy in quantifying miRNAs compared to the gold-standard ligation-based kit.
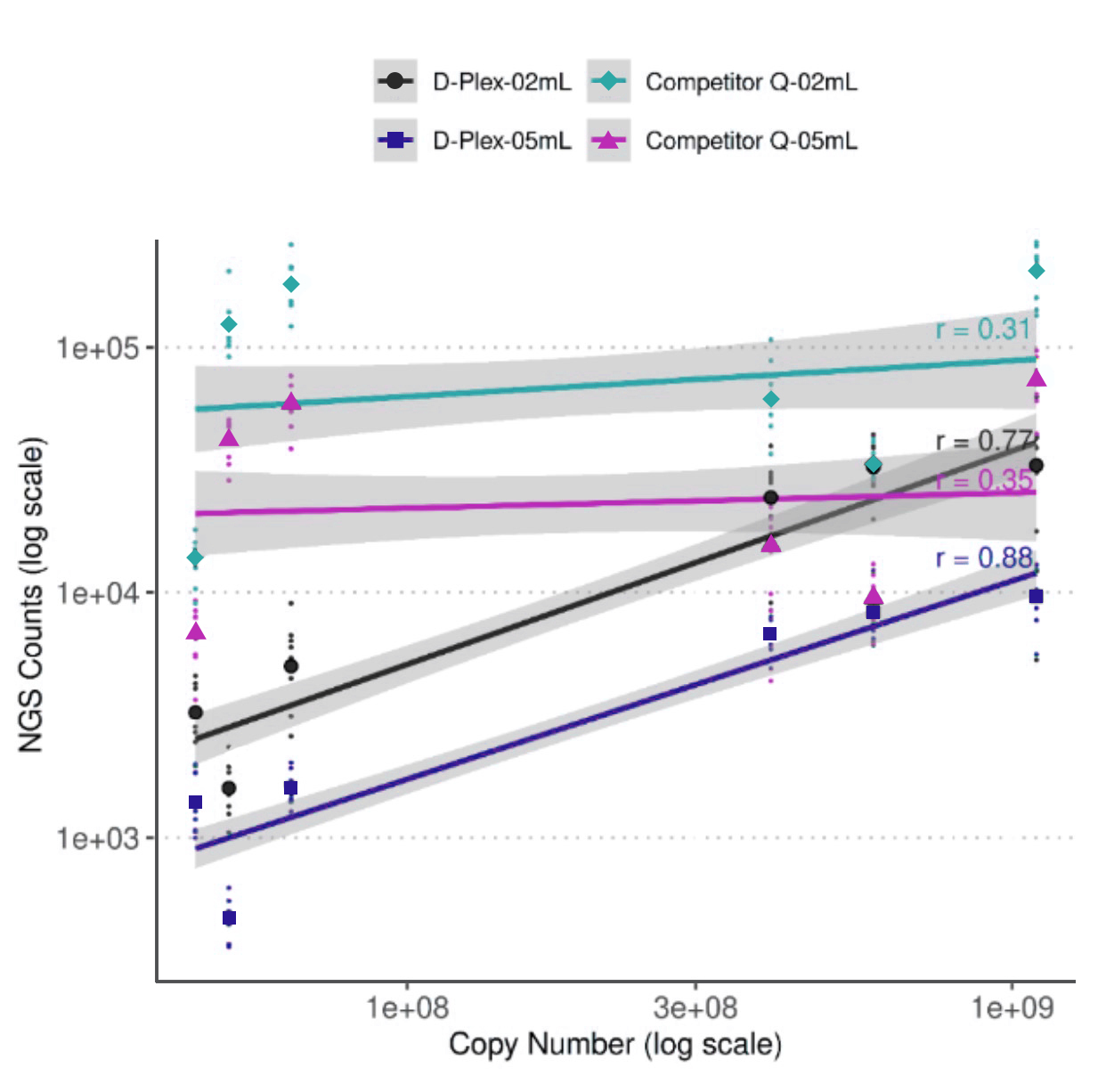
Figure 7. Six exogenous miRNA sequences were spiked into plasma in known quantities to measure the library quantification accuracy. The D-Plex cfRNA-seq maintains a linear relationship between NGS counts and spiked miRNA copy numbers, unlike competitor Q, which fails to show a proportional response. A high Pearson correlation value for D-Plex confirms its superior quantification accuracy.