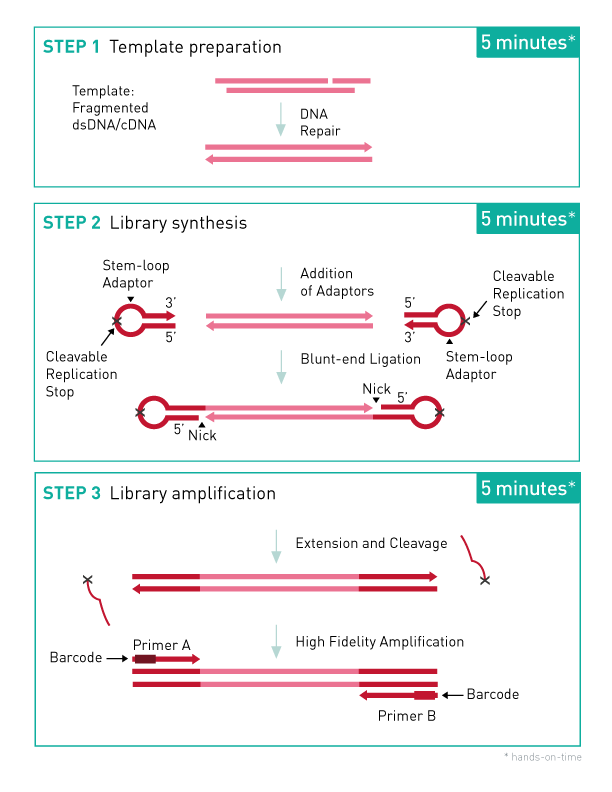
How to properly cite our product/service in your work We strongly recommend using this: MicroPlex Library Preparation Kit v3 /48 rxns (Hologic Diagenode Cat# C05010001). Click here to copy to clipboard. Using our products or services in your publication? Let us know! |
Comparative multi-omics of the macrophage response to infection with Mycobacterium tuberculosis complex bacteria reveals pathogen-driven epigenomic reprogramming
Hall, Thomas J.et al.
Background Bovine tuberculosis (bTB) is a chronic infectious disease primarily caused by Mycobacterium bovis, which inflicts significant economic losses on the global livestock industry worldwide and can also cause tuberculosis (TB) disease in other mammalian species, including humans. Alveolar macrophag... |
Acute NIPBL depletion reveals in vivo dynamics of loop extrusion and its role in transcription activation
Popay, Tessa M et al.
The organization of the genome in three-dimensional space is highly dynamic, yet how these dynamics are regulated and the role they play in genome function is poorly understood. Here we utilized acute depletion of NIPBL to characterize cohesin-mediated loop extrusion in vivo. We find that many chromatin loops ar... |
Multimodal epigenetic and enhancer network remodeling shape the transcriptional landscape of human beige adipocytes
Hazell Pickering, Sarah et al.
Epigenetic regulation is a key determinant of adipocyte fate, driving the differentiation toward white or thermogenic beige phenotypes in response to environmental cues. To dissect the mechanisms orchestrating this plasticity in human adipocytes, we conducted an integrative analysis of transcriptomic, epigenomic... |
Multimodal epigenetic and enhancer network remodeling shape the transcriptional landscape of human beige adipocytes
Hazell Pickering, Sarah et al.
Abstract
Epigenetic regulation is a key determinant of adipocyte fate, driving the differentiation toward white or thermogenic beige phenotypes in response to environmental cues. To dissect the mechanisms orchestrating this plasticity in human adipocytes, we conducted an integrative analysis of transcriptomic, epig... |
Epigenetic control of microglial developmental milestones from proliferative progenitors to efficient phagocytes
Pereira-Iglesias, Marta et al.
Early immune perturbations increase the risk of neurodegenerative and neurodevelopmental disorders, yet the mechanisms underlying the maturation of microglia, the resident immune cells of the brain parenchyma, remain poorly defined. Specifically, how proliferation, morphological differentiation, and phagocytosis are... |
Upregulation of ALDH1 as an adaptive epigenetic response to anthracyclines in acute myeloid leukemia
Leonetti, Francesco et al.
Acute myeloid leukemia (AML) is a genetically heterogeneous malignancy characterized by the clonal proliferation of undifferentiated myeloid precursors in the bone marrow. Although standard induction regimens based on anthracyclines often achieve initial remission, up to 25% of patients exhibit primary refractor... |
PHF19 drives the formation of PRC2 clusters to enhance motility in TNBC cells
Pelzer, Nina et al.
Polycomb repressive complex 2 (PRC2) is a key regulator of transcriptional repression and chromatin organization, essential in development and disease. While its enzymatic activity is well characterized, the factors governing PRC2 subnuclear organization, particularly in cancer cells, are largely unknown. Here, we... |
Gene and transposable element expression in response to stress in temperate and tropical populations of Drosophila
Merenciano, Miriam et al.
Background: The study of stress response in natural populations is crucial for understanding species local adaptation and evolution. In Drosophila, significant genetic diversity across populations from different geographical origins has been observed, emphasizing the influence of local environments.
Results:&n... |
Active chromatin marks and up-regulation of FOXC1 in uterine epithelial cells demarcate the onset of reproductive decline in aging females
Tsolova,Aleksandra O et al.
Advanced maternal age increases the risk of pregnancy complications due, in part, to changes in the uterine environment. Here, we show that uterine aging in mice is associated with a progressive increase in transcriptional variation, accompanied by a notable accumulation of activating histone marks at multiple gen... |
H1.3 depletion in AML cells prompts H1.2 redistribution, chromatin remodeling and cell cycle defects
Tellez-Quijorna, Clara et al.
Linker histone H1 variants play critical, yet distinct, roles in chromatin organization and gene regulation. However, very little is known about their specificity in cancer cells. In this study, we investigated the specificity of the H1.3 variant in acute myeloid leukemia (AML) cells. Through chromatin mapping and... |
SUMO operates from a unique long tandem repeat to keep innate immunity in check
Goffeney, Amandine et al.
The SUMO pathway mainly functions to repress innate immunity in myeloid cells. Inactivating sumoylation triggers a strong, noncanonical, type I interferon (IFN1) response, amplified and coupled with inflammation upon stimulation. These findings transposed to pre-clinical models with the demonstration that sumoyl... |
Shift in the urinary metabolome associated with 2,3,7,8-tetrachlorodibenzo-p-dioxin activation of the hepatic aryl hydrocarbon receptor
Sink, Warren J et al.
Epidemiological evidence suggests an association between dioxin and dioxin-like compound (DLC) exposure and human liver disease. In rodents, the prototypical DLC, 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD), has been shown to induce the progression of reversible hepatic steatosis to steatohepatitis with periporta... |
High-resolution map of the Plasmodium falciparum genome reveals MORC/ApiAP2-mediated links between distant, functionally related genes
Singh, Parul et al.
Genome organization plays an important role in silencing compacted, heterochromatinized genes in the most virulent human malaria parasite, Plasmodium falciparum. However, it remains unclear how these genes spatially cluster or whether active genes are also organized in a specific manner. We used Micro-C to achie... |
TP53 missense-specific transcriptional plasticity drives resistance against cell cycle inhibitors in pancreatic cancer
Urbach, Laura et al.
In ~70% of patients with pancreatic ductal adenocarcinoma, the TP53 gene acquires gain-of-function (GOF) mutations leading to rapid disease progression. Specifically, missense p53 (misp53) GOF mutations associate with therapy resistance and worse clinical outcomes. However, the molecular functions of dis... |
Overlapping and distinct functions of SPT6, PNUTS, and PCF11 in regulating transcription termination
Fabienne Bejjani et al.
The histone chaperone and transcription elongation factor SPT6 is integral to RNA polymerase II (RNAPII) activity. SPT6 also plays a crucial role in regulating transcription termination, although the mechanisms involved are largely unknown. In an attempt to identify the pathways employed by SPT6 in this regulation, ... |
TGIF2 is a major regulator of neural stem cell fate and neurogenic priming
Yiling Li et al.
During brain development, neural stem cells (NSCs) must balance self-renewal with differentiation and ensure lineage progression. To identify novel regulators of NSCs during neurogenesis, we isolated NSCs by FACS from the mouse cerebral cortex and ganglionic eminence at mid-neurogenesis, and at birth, when gliogenes... |
Accelerated epigenetic aging in Huntington’s disease involves polycomb repressive complex 1
Baptiste Brulé et al.
Loss of epigenetic information during physiological aging compromises cellular identity, leading to de-repression of developmental genes. Here, we assessed the epigenomic landscape of vulnerable neurons in two reference mouse models of Huntington neurodegenerative disease (HD), using cell-type-specific multi-omics, ... |
Ectopic expression of DNMT3L in human trophoblast stem cells restores features of the placental methylome
Lea, Georgia et al.
The placental DNA methylation landscape is unique, with widespread partially methylated domains (PMDs). The placental "methylome" is conserved across mammals, a shared feature of many cancers, and extensively studied for links with pregnancy complications. Human trophoblast stem cells (hTSCs) offer exciting potent... |
Effective in vivo binding energy landscape illustrates kinetic stability of RBPJ-DNA binding
Duyen Huynh et al.
Transcription factors (TFs) such as RBPJ in Notch signaling bind to specific DNA sequences to regulate transcription. How TF-DNA binding kinetics and cofactor interactions modulate gene regulation is mostly unknown. We determine the binding kinetics, transcriptional activity, and genome-wide chromatin occupation of ... |
Chromatin environment-dependent effects of DOT1L on gene expression in male germ cells
Manon Coulée et al.
The H3K79 methyltransferase DOT1L is essential for multiple aspects of mammalian development where it has been shown to regulate gene expression. Here, by producing and integrating epigenomic and spike-in RNA-seq data, we decipher the molecular role of DOT1L during mouse spermatogenesis and show that it has opposite... |
Trithorax regulates long-term memory in Drosophila through epigenetic maintenance of mushroom body metabolic state and translation capacity
Nicholas Raun et al.
The role of epigenetics and chromatin in the maintenance of postmitotic neuronal cell identities is not well understood. Here, we show that the histone methyltransferase Trithorax (Trx) is required in postmitotic memory neurons of the Drosophila mushroom body (MB) to enable their capacity for long-term mem... |
MECOM is a master repressor of myeloid differentiation through dose control of CEBPA in acute myeloid leukemia
Dorien Pastoors et al.
Enhancer translocations, due to 3q26 rearrangements, drive out-of-context MECOM expression in an aggressive subtype of acute myeloid leukemia (AML). Direct depletion of MECOM using an endogenous auxin-inducible degron immediately upregulates expression of myeloid differentiation factor CEBPA. MECOM depletion is also... |
RNA Pol-II transcripts in nucleolar associated domains of cancer cell nucleoli
Soumya Roy Chowdhury et al.
We performed a comparative study of the non-ribosomal gene content of nucleoli from seven cancer cell lines, using identical methods of purification and analysis. We identified unique chromosomal domains associated with the nucleolus (NADs) and genes within these domains (NAGs). Four cell lines have relatively few N... |
Interferon-gamma rescues FK506 dampened dendritic cell calcineurin-dependent responses to Aspergillus fumigatus via Stat3 to Stat1 switching
Amit Adlakha et al.
IScience Highlights
Calcineurin inhibitors block DC maturation in response to A. fumigatus
Lack of DC maturation impairs Th1 polarization in response to A. fumigatus
Interferon-γ restores maturation, promotes Th1 polarization and fungal killing
ChIPseq reveals in... |
ETV2/ER71 regulates hematovascular lineage generation and vascularization through an H3K9 demethylase, KDM4A
Min Seong Kim et al.
Highlights
Interaction of ETV2 and KDM4A decreases H3K9 trimethylation on hematovascular genes.
ETV2 and KDM4A cooperatively regulates the expression of hematovascular genes.
Mice lacking endothelial Etv2 and Kdm4a display a severe angiogenic impairment.
... |
Epi-microRNA mediated metabolic reprogramming counteracts hypoxia to preserve affinity maturation
Rinako Nakagawa et al.
To increase antibody affinity against pathogens, positively selected GC-B cells initiate cell division in the light zone (LZ) of germinal centers (GCs). Among these, higher-affinity clones migrate to the dark zone (DZ) and vigorously proliferate by utilizing energy provided by oxidative phosphorylation (OXPHOS). How... |
TEAD-targeting small molecules induce a cofactor switch to regulate the Hippo pathway
Alissa D. Guarnaccia et al.
TEAD proteins are the main transcriptional effectors of the Hippo signaling pathway and a pharmacological target in oncology. Most TEAD-targeting small molecules act by disrupting interaction with the oncogenic transcriptional activators YAP and TAZ. Here, we describe an alternative mechanism for TEAD lipid pocket b... |
ACE2 Enhances Sensitivity to PD-L1 Blockade by Inhibiting Macrophage-Induced Immunosuppression and Angiogenesis
Peiyi Xie et al.
Anti-PD-L1-based combination immunotherapy has become the first-line treatment for unresectable hepatocellular carcinoma (HCC). However, the objective response rate is lower than 40%, highlighting the need to identify mechanisms of tolerance to immune checkpoint inhibitors and accurate biomarkers of response. Here, ... |
Extracellular vesicle-mediated trafficking of molecular cues during human brain development
Forero A. et al.
Cellular crosstalk is an essential process influenced by numerous factors, including secreted vesicles that transfer nucleic acids, lipids, and proteins between cells. Extracellular vesicles (EVs) have been the center of many studies focusing on neurodegenerative disorders, but whether EVs display cell-type-specific... |
The small inhibitor WM-1119 effectively targets KAT6A-rearranged AML, but not KMT2A-rearranged AML, despite shared KAT6 genetic dependency
Mathew Sheridan et al.
Background
The epigenetic factors KAT6A (MOZ/MYST3) and KMT2A (MLL/MLL1) interact in normal hematopoiesis to regulate progenitors’ self-renewal. Both proteins are recurrently translocated in AML, leading to impairment of critical differentiation pathways in these malignant cells. We evaluated the potential of... |
Integrated multi-omics analysis of PBX1 in mouse adult neural stem- and progenitor cells identifies a transcriptional module that functionally links PBX1 to TCF3/4
Vera Laub et al.
Developmental transcription factors act in networks, but how these networks achieve cell- and tissue specificity is still poorly understood. Here, we explored pre-B cell leukemia homeobox 1 (PBX1) in adult neurogenesis combining genomic, transcriptomic, and proteomic approaches. ChIP-seq analysis uncovered PBX1 bind... |
HNF1β bookmarking involves Topoisomerase 1 activation and DNA topology relaxation in mitotic chromatin
Alessia Bagattin et al.
Highlights
HNF1β mitotic site binding is preserved with a specific methanol/formaldehyde ChIP
BTBD2, an HNF1β partner, mediates mitosis-specific interaction with TOP1
HNF1β recruits TOP1 and induces DNA relaxation around bookmarked HNF1β sites
An HNF1β m... |
RNA polymerase II transcription initiation in holo-TFIID-depleted mouse embryonic stem cells
Hisler V. et al.
The recognition of core promoter sequences by TFIID is the first step in RNA polymerase II (Pol II) transcription initiation. Metazoan holo-TFIID is a trilobular complex, composed of the TATA binding protein (TBP) and 13 TBP-associated factors (TAFs). Why and how TAFs are necessary for the formation of TFIID domains... |
An atlas of the human liver diurnal transcriptome and its perturbation by hepatitis C virus infection
Mukherji A. et al.
Chronic liver disease and cancer are global health challenges. The role of the circadian clock as a regulator of liver physiology and disease is well established in rodents, however, the identity and epigenetic regulation of rhythmically expressed genes in human disease is less well studied. Here we unravel the rhyt... |
Innate immune training restores pro-reparative myeloid functions to promote remyelination in the aged central nervous system
Tiwari V. et al.
The reduced ability of the central nervous system to regenerate with increasing age limits functional recovery following demyelinating injury. Previous work has shown that myelin debris can overwhelm the metabolic capacity of microglia, thereby impeding tissue regeneration in aging, but the underlying mechanisms are... |
A multiomic atlas of the aging hippocampus reveals molecular changes in response to environmental enrichment
Perez R. F. at al.
Aging involves the deterioration of organismal function, leading to the emergence of multiple pathologies. Environmental stimuli, including lifestyle, can influence the trajectory of this process and may be used as tools in the pursuit of healthy aging. To evaluate the role of epigenetic mechanisms in this context, ... |
Epigenetic alterations affecting hematopoietic regulatory networks as drivers of mixed myeloid/lymphoid leukemia
Roger Mulet-Lazaro et al.
Leukemias with ambiguous lineage comprise several loosely defined entities, often without a clear mechanistic basis. Here, we extensively profile the epigenome and transcriptome of a subgroup of such leukemias with CpG Island Methylator Phenotype. These leukemias exhibit comparable hybrid myeloid/lymphoid epigenetic... |
Multiomics uncovers the epigenomic and transcriptomic response to viral and bacterial stimulation in turbot
Aramburu O. et al.
Uncovering the epigenomic regulation of immune responses is essential for a comprehensive understanding of host defence mechanisms but remains poorly described in farmed fish. Here, we report the first annotation of the innate immune regulatory response in the genome of turbot (Scophthalmus maximus), a farmed flatfi... |
SMARCB1 loss activates patient-specific distal oncogenic enhancers in malignant rhabdoid tumors
Liu, N.Q. et al.
Malignant rhabdoid tumor (MRT) is a highly malignant and often lethal childhood cancer. MRTs are genetically defined by bi-allelic inactivating mutations in SMARCB1, a member of the BRG1/BRM-associated factors (BAF) chromatin remodeling complex. Mutations in BAF complex members are common in human cancer, yet t... |
The genetic landscape of origins of replication in P. falciparum
Casilda Muñoz Castellano et al.
Various origin mapping approaches have enabled genome-wide identification of origins of replication (ORI) in model organisms, but only a few studies have focused on divergent organisms. By employing three complementary approaches we provide a high-resolution map of ORIs in Plasmodium falciparum, the deadliest h... |
Therapeutic targeting of EP300/CBP by bromodomain inhibition in hematologic malignancies
Luciano Nicosia et al.
CCS1477 (inobrodib) is a potent, selective EP300/CBP bromodomain inhibitor which induces cell-cycle arrest and differentiation in hematologic malignancy model systems. In myeloid leukemia cells, it promotes rapid eviction of EP300/CBP from an enhancer subset marked by strong MYB occupancy and high H3K27 acetylation,... |
Differentiation block in acute myeloid leukemia regulated by intronicsequences of FTO
Camera F. et al.
Iroquois transcription factor gene IRX3 is highly expressed in 20–30\% of acute myeloid leukemia (AML) and contributes to the pathognomonic differentiation block. Intron 8 FTO sequences ∼220kB downstream of IRX3 exhibit histone acetylation, DNA methylation, and contacts with th... |
Supraphysiological Androgens Promote the Tumor Suppressive Activity of the Androgen Receptor Through cMYC Repression and Recruitment of the DREAM Complex
Nyquist M. et al.
The androgen receptor (AR) pathway regulates key cell survival programs in prostate epithelium. The AR represents a near-universal driver and therapeutic vulnerability in metastatic prostate cancer, and targeting AR has a remarkable therapeutic index. Though most approaches directed toward AR focus on inhibiting AR ... |
In skeletal muscle and neural crest cells, SMCHD1 regulates biologicalpathways relevant for Bosma syndrome and facioscapulohumeral dystrophyphenotype.
Laberthonnière C. et al.
Many genetic syndromes are linked to mutations in genes encoding factors that guide chromatin organization. Among them, several distinct rare genetic diseases are linked to mutations in SMCHD1 that encodes the structural maintenance of chromosomes flexible hinge domain containing 1 chromatin-associated factor. In hu... |
Hypomethylation and overexpression of Th17-associated genes is ahallmark of intestinal CD4+ lymphocytes in Crohn's disease.
Sun Z. et al.
BACKGROUND: The development of Crohn's disease (CD) involves immune cell signaling pathways regulated by epigenetic modifications. Aberrant DNA methylation has been identified in peripheral blood and bulk intestinal tissue from CD patients. However, the DNA methylome of disease-associated intestinal CD4 + lymphocyte... |
Mutant FUS induces chromatin reorganization in the hippocampus andalters memory processes.
Tzeplaeff L. et al.
Cytoplasmic mislocalization of the nuclear Fused in Sarcoma (FUS) protein is associated to amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). Cytoplasmic FUS accumulation is recapitulated in the frontal cortex and spinal cord of heterozygous Fus mice. Yet, the mechanisms linking FUS mislocalizati... |
Transient suppression of SUMOylation in embryonic stem cells generatesembryo-like structures.
Cossec J-C. et al.
Recent advances in synthetic embryology have opened new avenues for understanding the complex events controlling mammalian peri-implantation development. Here, we show that mouse embryonic stem cells (ESCs) solely exposed to chemical inhibition of SUMOylation generate embryo-like structures comprising anterior neura... |
ZEB1 controls a lineage-specific transcriptional program essential formelanoma cell state transitions
Tang Y. et al.
Cell plasticity sustains intra-tumor heterogeneity and treatment resistance in melanoma. Deciphering the transcriptional mechanisms governing reversible phenotypic transitions between proliferative/differentiated and invasive/stem-like states is required in order to design novel therapeutic strategies. EMT-inducing ... |
Impact of Fetal Exposure to Endocrine Disrupting ChemicalMixtures on FOXA3 Gene and Protein Expression in Adult RatTestes.
Walker C. et al.
Perinatal exposure to endocrine disrupting chemicals (EDCs) has been shown to affect male reproductive functions. However, the effects on male reproduction of exposure to EDC mixtures at doses relevant to humans have not been fully characterized. In previous studies, we found that in utero exposure to mixtures of th... |
TCDD induces multigenerational alterations in the expression ofmicroRNA in the thymus through epigenetic modifications
Singh Narendra P et al.
2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD), a potent AhR ligand, is an environmental contaminant that is known for mediating toxicity across generations. However, whether TCDD can induce multigenerational changes in the expression of miRNAs (miRs) has not been previously studied. In the current study, we investigate... |
The histone acetyltransferase KAT6A is recruited to unmethylatedCpG islands via a DNA binding winged helix domain.
Weber L.M. et al.
The lysine acetyltransferase KAT6A (MOZ, MYST3) belongs to the MYST family of chromatin regulators, facilitating histone acetylation. Dysregulation of KAT6A has been implicated in developmental syndromes and the onset of acute myeloid leukemia (AML). Previous work suggests that KAT6A is recruited to its genomic targ... |
Polyglutamine-expanded ATXN7 alters a specific epigenetic signatureunderlying photoreceptor identity gene expression in SCA7 mouseretinopathy.
Niewiadomska-Cimicka A.et al.
BACKGROUND: Spinocerebellar ataxia type 7 (SCA7) is a neurodegenerative disorder that primarily affects the cerebellum and retina. SCA7 is caused by a polyglutamine expansion in the ATXN7 protein, a subunit of the transcriptional coactivator SAGA that acetylates histone H3 to deposit narrow H3K9ac mark at DNA regula... |
Intranasal administration of Acinetobacter lwoffii in a murine model ofasthma induces IL-6-mediated protection associated with cecal microbiotachanges.
Alashkar A. B. et al.
BACKGROUND: Early-life exposure to certain environmental bacteria including Acinetobacter lwoffii (AL) has been implicated in protection from chronic inflammatory diseases including asthma later in life. However, the underlying mechanisms at the immune-microbe interface remain largely unknown. METHODS: The effects o... |
Cryptococcal Hsf3 controls intramitochondrial ROS homeostasis byregulating the respiratory process.
Gao X.et al.
Mitochondrial quality control prevents accumulation of intramitochondrial-derived reactive oxygen species (mtROS), thereby protecting cells against DNA damage, genome instability, and programmed cell death. However, underlying mechanisms are incompletely understood, particularly in fungal species. Here, we show that... |
Dominant role of DNA methylation over H3K9me3 for IAP silencingin endoderm.
Wang Z. et al.
Silencing of endogenous retroviruses (ERVs) is largely mediated by repressive chromatin modifications H3K9me3 and DNA methylation. On ERVs, these modifications are mainly deposited by the histone methyltransferase Setdb1 and by the maintenance DNA methyltransferase Dnmt1. Knock-out of either Setdb1 or Dnmt1 leads to... |
HDAC1 and PRC2 mediate combinatorial control in SPI1/PU.1-dependentgene repression in murine erythroleukaemia.
Gregoricchio S. et al.
Although originally described as transcriptional activator, SPI1/PU.1, a major player in haematopoiesis whose alterations are associated with haematological malignancies, has the ability to repress transcription. Here, we investigated the mechanisms underlying gene repression in the erythroid lineage, in which SPI1 ... |
Dual role of histone variant H3.3B in spermatogenesis: positiveregulation of piRNA transcription and implication in X-chromosomeinactivation.
Fontaine E. et al.
The histone variant H3.3 is encoded by two distinct genes, H3f3a and H3f3b, exhibiting identical amino-acid sequence. H3.3 is required for spermatogenesis, but the molecular mechanism of its spermatogenic function remains obscure. Here, we have studied the role of each one of H3.3A and H3.3B proteins in spermatogene... |
TBX2 acts as a potent transcriptional silencer of tumour suppressor genesthrough interaction with the CoREST complex to sustain theproliferation of breast cancers.
McIntyre A.J. et al.
Chromosome 17q23 amplification occurs in 20\% of primary breast tumours and is associated with poor outcome. The TBX2 gene is located on 17q23 and is often over-expressed in this breast tumour subset. TBX2 is an anti-senescence gene, promoting cell growth and survival through repression of Tumour Suppressor Genes (T... |
Caffeine intake exerts dual genome-wide effects on hippocampal metabolismand learning-dependent transcription.
Paiva I. et al.
Caffeine is the most widely consumed psychoactive substance in the world. Strikingly, the molecular pathways engaged by its regular consumption remain unclear. We herein addressed the mechanisms associated with habitual (chronic) caffeine consumption in the mouse hippocampus using untargeted orthogonal omics techniq... |
Epigenetic Mechanisms Mediating Cell State Transitions in Chondrocytes
Wuelling M. et al.
Epigenetic modifications play critical roles in regulating cell lineage differentiation, but the epigenetic mechanisms guiding specific differentiation steps within a cell lineage have rarely been investigated. To decipher such mechanisms, we used the defined transition from proliferating (PC) into hypertrophic chon... |
The CpG Island-Binding Protein SAMD1 Contributes to anUnfavorable Gene Signature in HepG2 Hepatocellular CarcinomaCells.
Simon C. et al.
The unmethylated CpG island-binding protein SAMD1 is upregulated in many human cancer types, but its cancer-related role has not yet been investigated. Here, we used the hepatocellular carcinoma cell line HepG2 as a cancer model and investigated the cellular and transcriptional roles of SAMD1 using ChIP-Seq and RNA-... |
Local euchromatin enrichment in lamina-associated domains anticipatestheir repositioning in the adipogenic lineage.
Madsen-Østerbye J. et al.
BACKGROUND: Interactions of chromatin with the nuclear lamina via lamina-associated domains (LADs) confer structural stability to the genome. The dynamics of positioning of LADs during differentiation, and how LADs impinge on developmental gene expression, remains, however, elusive. RESULTS: We examined changes in t... |
NuA4 and H2A.Z control environmental responses and autotrophicgrowth in Arabidopsis
Bieluszewski T. et al.
Nucleosomal acetyltransferase of H4 (NuA4) is an essential transcriptional coactivator in eukaryotes, but remains poorly characterized in plants. Here, we describe Arabidopsis homologs of the NuA4 scaffold proteins Enhancer of Polycomb-Like 1 (AtEPL1) and Esa1-Associated Factor 1 (AtEAF1). Loss of AtEAF1 results in ... |
Three classes of epigenomic regulators converge to hyperactivate theessential maternal gene deadhead within a heterochromatin mini-domain.
Torres-Campana D. et al.
The formation of a diploid zygote is a highly complex cellular process that is entirely controlled by maternal gene products stored in the egg cytoplasm. This highly specialized transcriptional program is tightly controlled at the chromatin level in the female germline. As an extreme case in point, the massive and s... |
Epromoters function as a hub to recruit key transcription factorsrequired for the inflammatory response
Santiago-Algarra D. et al.
Gene expression is controlled by the involvement of gene-proximal (promoters) and distal (enhancers) regulatory elements. Our previous results demonstrated that a subset of gene promoters, termed Epromoters, work as bona fide enhancers and regulate distal gene expression. Here, we hypothesized that Epromoters play a... |
Decreased PRC2 activity supports the survival of basal-like breastcancer cells to cytotoxic treatments
Mieczkowska IK et al.
Breast cancer (BC) is the most common cancer occurring in women but also rarely develops in men. Recent advances in early diagnosis and development of targeted therapies have greatly improved the survival rate of BC patients. However, the basal-like BC subtype (BLBC), largely overlapping with the triple-negative BC ... |
Ago1 controls myogenic differentiation by regulating eRNA-mediatedCBP-guided epigenome reprogramming.
Fallatah Bodor et al.
The role of chromatin-associated RNAi components in the nucleus of mammalian cells and in particular in the context of developmental programs remains to be elucidated. Here, we investigate the function of nuclear Argonaute 1 (Ago1) in gene expression regulation during skeletal muscle differentiation. We show that Ag... |
Extensive NEUROG3 occupancy in the human pancreatic endocrine generegulatory network.
Schreiber V. et al.
OBJECTIVE: Mice lacking the bHLH transcription factor (TF) Neurog3 do not form pancreatic islet cells, including insulin-secreting beta cells, the absence of which leads to diabetes. In humans, homozygous mutations of NEUROG3 manifest with neonatal or childhood diabetes. Despite this critical role in islet cell deve... |
Alveolar macrophages from persons living with HIV show impairedepigenetic response to Mycobacterium tuberculosis.
Correa-Macedo Wilian et al.
Persons living with HIV (PLWH) are at increased risk of tuberculosis (TB). HIV-associated TB is often the result of recent infection with Mycobacterium tuberculosis (Mtb) followed by rapid progression to disease. Alveolar macrophages (AM) are the first cells of the innate immune system that engage Mtb, but how HIV a... |
INTS11 regulates hematopoiesis by promoting PRC2 function.
Zhang Peng et al.
INTS11, the catalytic subunit of the Integrator (INT) complex, is crucial for the biogenesis of small nuclear RNAs and enhancer RNAs. However, the role of INTS11 in hematopoietic stem and progenitor cell (HSPC) biology is unknown. Here, we report that INTS11 is required for normal hematopoiesis and hematopoietic-spe... |
The related coactivator complexes SAGA and ATAC control embryonicstem cell self-renewal through acetyltransferase-independent mechanisms
Fischer Veronique et al.
SUMMARY SAGA (Spt-Ada-Gcn5 acetyltransferase) and ATAC (Ada-two-A-containing) are two related coactivator complexes, sharing the same histone acetyltransferase (HAT) subunit. The HAT activities of SAGA and ATAC are required for metazoan development, but the role of these complexes in RNA polymerase II transcription ... |
Altered Chromatin States Drive Cryptic Transcription in AgingMammalian Stem Cells.
McCauley Brenna S et al.
A repressive chromatin state featuring trimethylated lysine 36 on histone H3 (H3K36me3) and DNA methylation suppresses cryptic transcription in embryonic stem cells. Cryptic transcription is elevated with age in yeast and nematodes, and reducing it extends yeast lifespan, though whether this occurs in mammals is unk... |
Metabolically controlled histone H4K5 acylation/acetylation ratiodrives BRD4 genomic distribution.
Gao M. et al.
In addition to acetylation, histones are modified by a series of competing longer-chain acylations. Most of these acylation marks are enriched and co-exist with acetylation on active gene regulatory elements. Their seemingly redundant functions hinder our understanding of histone acylations' specific roles. Here, by... |
The SAM domain-containing protein 1 (SAMD1) acts as a repressivechromatin regulator at unmethylated CpG islands
Stielow B. et al.
CpG islands (CGIs) are key regulatory DNA elements at most promoters, but how they influence the chromatin status and transcription remains elusive. Here, we identify and characterize SAMD1 (SAM domain-containing protein 1) as an unmethylated CGI-binding protein. SAMD1 has an atypical winged-helix domain that direct... |
Polycomb Repressive Complex 2 and KRYPTONITE regulate pathogen-inducedprogrammed cell death in Arabidopsis.
Dvořák Tomaštíková E. et al.
The Polycomb Repressive Complex 2 (PRC2) is well-known for its role in controlling developmental transitions by suppressing the premature expression of key developmental regulators. Previous work revealed that PRC2 also controls the onset of senescence, a form of developmental programmed cell death (PCD) in plants. ... |
An integrated multi-omics analysis identifies prognostic molecularsubtypes of non-muscle-invasive bladder cancer
Lindskrog Sia Viborg et al.
The molecular landscape in non-muscle-invasive bladder cancer (NMIBC) is characterized by large biological heterogeneity with variable clinical outcomes. Here, we perform an integrative multi-omics analysis of patients diagnosed with NMIBC (n = 834). Transcriptomic analysis identifies four classes (1, ... |
VPRBP functions downstream of the androgen receptor and OGT to restrict p53 activation in prostate cancer
Poulose N. et al.
Androgen receptor (AR) is a major driver of prostate cancer (PCa) initiation and progression. O-GlcNAc transferase (OGT), the enzyme that catalyses the covalent addition of UDP-N-acetylglucosamine (UDP-GlcNAc) to serine and threonine residues of proteins, is often up-regulated in PCa with its expression correlated w... |
JAZF1, A Novel p400/TIP60/NuA4 Complex Member, Regulates H2A.ZAcetylation at Regulatory Regions.
Procida, Tara and Friedrich, Tobias and Jack, Antonia P M and Peritore,Martina and Bönisch, Clemens and Eberl, H Christian and Daus, Nadine andKletenkov, Konstantin and Nist, Andrea and Stiewe, Thorsten and Borggrefe,Tilman and Mann, Matthias and Bartk
Histone variants differ in amino acid sequence, expression timing and genomic localization sites from canonical histones and convey unique functions to eukaryotic cells. Their tightly controlled spatial and temporal deposition into specific chromatin regions is accomplished by dedicated chaperone and/or remodeling c... |
Histone demethylase JMJD2B/KDM4B regulates transcriptional program viadistinctive epigenetic targets and protein interactors for the maintenanceof trophoblast stem cells.
Mak, Kylie Hin-Man et al.
Trophoblast stem cell (TSC) is crucial to the formation of placenta in mammals. Histone demethylase JMJD2 (also known as KDM4) family proteins have been previously shown to support self-renewal and differentiation of stem cells. However, their roles in the context of the trophoblast lineage remain unclear. Here, we ... |
Trans- and cis-acting effects of Firre on epigenetic features of theinactive X chromosome.
Fang, He and Bonora, Giancarlo and Lewandowski, Jordan P and Thakur,Jitendra and Filippova, Galina N and Henikoff, Steven and Shendure, Jay andDuan, Zhijun and Rinn, John L and Deng, Xinxian and Noble, William S andDisteche, Christine M
Firre encodes a lncRNA involved in nuclear organization. Here, we show that Firre RNA expressed from the active X chromosome maintains histone H3K27me3 enrichment on the inactive X chromosome (Xi) in somatic cells. This trans-acting effect involves SUZ12, reflecting interactions between Firre RNA and components of t... |
H2A.Z is dispensable for both basal and activated transcription inpost-mitotic mouse muscles.
Belotti E. et al.
While the histone variant H2A.Z is known to be required for mitosis, it is also enriched in nucleosomes surrounding the transcription start site of active promoters, implicating H2A.Z in transcription. However, evidence obtained so far mainly rely on correlational data generated in actively dividing cells. We have e... |